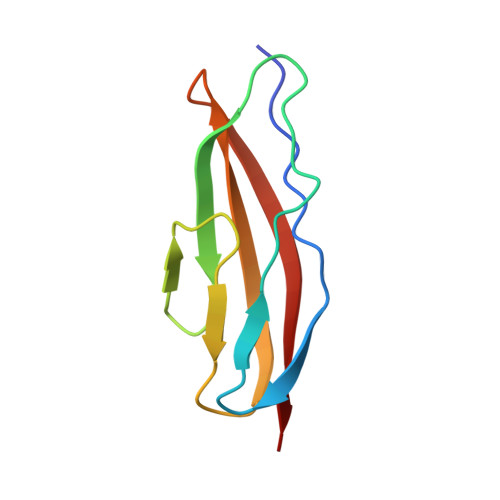
Structures of three polycystic kidney disease-like domains from Clostridium histolyticum collagenases ColG and ColH.
Bauer, R., Janowska, K., Taylor, K., Jordan, B., Gann, S., Janowski, T., Latimer, E.C., Matsushita, O., Sakon, J.(2015) Acta Crystallogr D Biol Crystallogr 71: 565-577
- PubMed: 25760606 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004714027722
- Primary Citation Related Structures:
4JGU, 4JRW, 4TN9, 4U6T, 4U7K - PubMed Abstract:
Clostridium histolyticum collagenases ColG and ColH are segmental enzymes that are thought to be activated by Ca(2+)-triggered domain reorientation to cause extensive tissue destruction. The collagenases consist of a collagenase module (s1), a variable number of polycystic kidney disease-like (PKD-like) domains (s2a and s2b in ColH and s2 in ColG) and a variable number of collagen-binding domains (s3 in ColH and s3a and s3b in ColG). The X-ray crystal structures of Ca(2+)-bound holo s2b (1.4 Å resolution, R = 15.0%, Rfree = 19.1%) and holo s2a (1.9 Å resolution, R = 16.3%, Rfree = 20.7%), as well as of Ca(2+)-free apo s2a (1.8 Å resolution, R = 20.7%, Rfree = 27.2%) and two new forms of N-terminally truncated apo s2 (1.4 Å resolution, R = 16.9%, Rfree = 21.2%; 1.6 Å resolution, R = 16.2%, Rfree = 19.2%), are reported. The structurally similar PKD-like domains resemble the V-set Ig fold. In addition to a conserved β-bulge, the PKD-like domains feature a second bulge that also changes the allegiance of the subsequent β-strand. This β-bulge and the genesis of a Ca(2+) pocket in the archaeal PKD-like domain suggest a close kinship between bacterial and archaeal PKD-like domains. Different surface properties and indications of different dynamics suggest unique roles for the PKD-like domains in ColG and in ColH. Surface aromatic residues found on ColH s2a-s2b, but not on ColG s2, may provide the weak interaction in the biphasic collagen-binding mode previously found in s2b-s3. B-factor analyses suggest that in the presence of Ca(2+) the midsection of s2 becomes more flexible but the midsections of s2a and s2b stay rigid. The different surface properties and dynamics of the domains suggest that the PKD-like domains of M9B bacterial collagenase can be grouped into either a ColG subset or a ColH subset. The conserved properties of PKD-like domains in ColG and in ColH include Ca(2+) binding. Conserved residues not only interact with Ca(2+), but also position the Ca(2+)-interacting water molecule. Ca(2+) aligns the N-terminal linker approximately parallel to the major axis of the domain. Ca(2+) binding also increases stability against heat and guanidine hydrochloride, and may improve the longevity in the extracellular matrix. The results of this study will further assist in developing collagen-targeting vehicles for various signal molecules.
- Department of Chemistry and Biochemistry, University of Arkansas, Fayetteville, AR 72701, USA.
Organizational Affiliation: