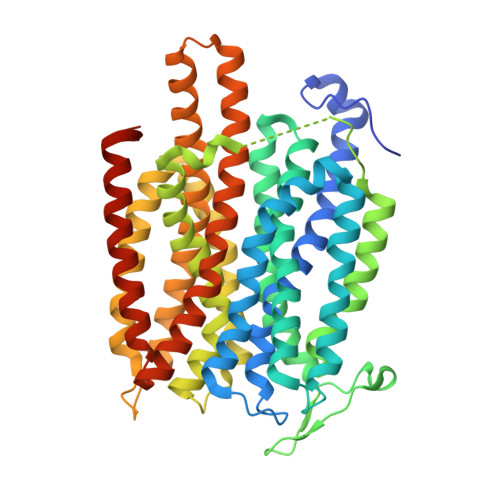
Structural basis for dynamic mechanism of nitrate/nitrite antiport by NarK
Fukuda, M., Takeda, H., Kato, H.E., Doki, S., Ito, K., Maturana, A.D., Ishitani, R., Nureki, O.(2015) Nat Commun 6: 7097-7097
- PubMed: 25959928 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms8097
- Primary Citation Related Structures:
4U4T, 4U4V, 4U4W - PubMed Abstract:
NarK belongs to the nitrate/nitrite porter (NNP) family in the major facilitator superfamily (MFS) and plays a central role in nitrate uptake across the membrane in diverse organisms, including archaea, bacteria, fungi and plants. Although previous studies provided insight into the overall structure and the substrate recognition of NarK, its molecular mechanism, including the driving force for nitrate transport, remained elusive. Here we demonstrate that NarK is a nitrate/nitrite antiporter, using an in vitro reconstituted system. Furthermore, we present the high-resolution crystal structures of NarK from Escherichia coli in the nitrate-bound occluded, nitrate-bound inward-open and apo inward-open states. The integrated structural, functional and computational analyses reveal the nitrate/nitrite antiport mechanism of NarK, in which substrate recognition is coupled to the transport cycle by the concomitant movement of the transmembrane helices and the key tyrosine and arginine residues in the substrate-binding site.
- 1] Department of Biological Sciences, Graduate School of Science, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan [2] Global Research Cluster, RIKEN, 2-1 Hirosawa, Wako-shi, Saitama 351-0198, Japan.
Organizational Affiliation: