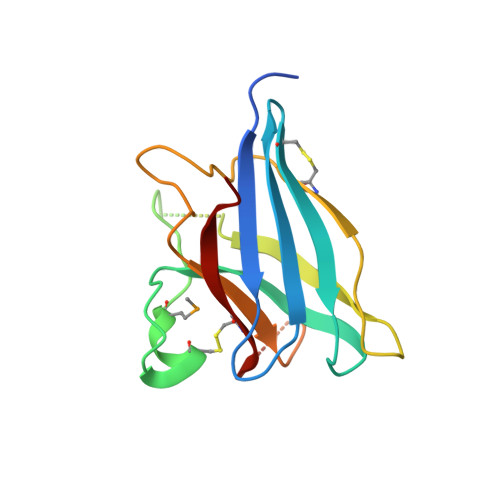
The megavirus chilensis cu,zn-superoxide dismutase: the first viral structure of a typical cellular copper chaperone-independent hyperstable dimeric enzyme.
Lartigue, A., Burlat, B., Coutard, B., Chaspoul, F., Claverie, J.M., Abergel, C.(2015) J Virol 89: 824-832
- PubMed: 25355875
- DOI: https://doi.org/10.1128/JVI.02588-14
- Primary Citation Related Structures:
4U4I - PubMed Abstract:
Giant viruses able to replicate in Acanthamoeba castellanii penetrate their host through phagocytosis. After capsid opening, a fusion between the internal membranes of the virion and the phagocytic vacuole triggers the transfer in the cytoplasm of the viral DNA together with the DNA repair enzymes and the transcription machinery present in the particles. In addition, the proteome analysis of purified mimivirus virions revealed the presence of many enzymes meant to resist oxidative stress and conserved in the Mimiviridae. Megavirus chilensis encodes a predicted copper, zinc superoxide dismutase (Cu,Zn-SOD), an enzyme known to detoxify reactive oxygen species released in the course of host defense reactions. While it was thought that the metal ions are required for the formation of the active-site lid and dimer stabilization, megavirus chilensis SOD forms a very stable metal-free dimer. We used electron paramagnetic resonance (EPR) analysis and activity measurements to show that the supplementation of the bacterial culture with copper and zinc during the recombinant expression of Mg277 is sufficient to restore a fully active holoenzyme. These results demonstrate that the viral enzyme's activation is independent of a chaperone both for disulfide bridge formation and for copper incorporation and suggest that its assembly may not be as regulated as that of its cellular counterparts. A SOD protein is encoded by a variety of DNA viruses but is absent from mimivirus. As in poxviruses, the enzyme might be dispensable when the virus infects Acanthamoeba cells but may allow megavirus chilensis to infect a broad range of eukaryotic hosts. Mimiviridae are giant viruses encoding more than 1,000 proteins. The virion particles are loaded with proteins used by the virus to resist the vacuole's oxidative stress. The megavirus chilensis virion contains a predicted copper, zinc superoxide dismutase (Cu,Zn-SOD). The corresponding gene is present in some megavirus chilensis relatives but is absent from mimivirus. This first crystallographic structure of a viral Cu,Zn-SOD highlights the features that it has in common with and its differences from cellular SODs. It corresponds to a very stable dimer of the apo form of the enzyme. We demonstrate that upon supplementation of the growth medium with Cu and Zn, the recombinant protein is fully active, suggesting that the virus's SOD activation is independent of a copper chaperone for SOD generally used by eukaryotic SODs.
- Structural and Genomic Information Laboratory, IGS UMR7256, Centre National de la Recherche Scientifique, Aix-Marseille Université, Institut de Microbiologie de la Méditerranée (IMM), Marseille, France.
Organizational Affiliation: