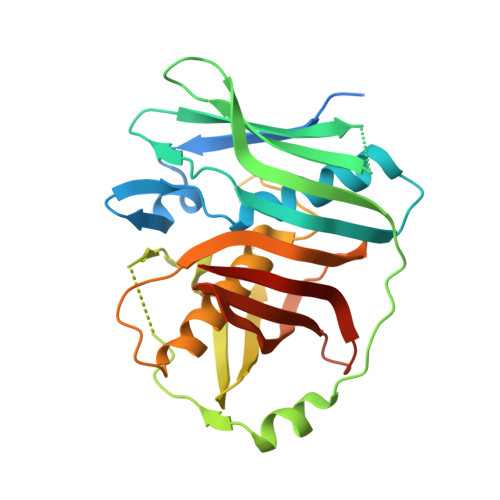
A Double-Hotdog with a New Trick: Structure and Mechanism of the trans-Acyltransferase Polyketide Synthase Enoyl-isomerase.
Gay, D.C., Spear, P.J., Keatinge-Clay, A.T.(2014) ACS Chem Biol 9: 2374-2381
- PubMed: 25089587 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/cb500459b
- Primary Citation Related Structures:
4U3V - PubMed Abstract:
Many polyketide natural products exhibit invaluable medicinal properties, yet much remains to be understood regarding the machinery responsible for their biosynthesis. The recently discovered trans-acyltransferase polyketide synthases employ processing enzymes that catalyze modifications unique from those of the classical cis-acyltransferase polyketide synthases. The enoyl-isomerase domains of these megasynthases shift double bonds and are well-represented by an enzyme that helps forge the triene system within the antibiotic produced by the prototypical bacillaene synthase. This first crystal structure of an enoyl-isomerase, at 1.73 Å resolution, not only revealed relationships between this class of enzymes and dehydratases but also guided an investigation into the mechanism of double bond migration. The catalytic histidine, positioned differently from that of dehydratases, was demonstrated to independently shuttle a proton between the γ- and α-positions of the intermediate. This unprecedented mechanism highlights the catalytic diversity of divergent enzymes within trans-acyltransferase polyketide synthases.
- Department of Molecular Biosciences, University of Texas at Austin , Austin, Texas 78712, United States.
Organizational Affiliation: