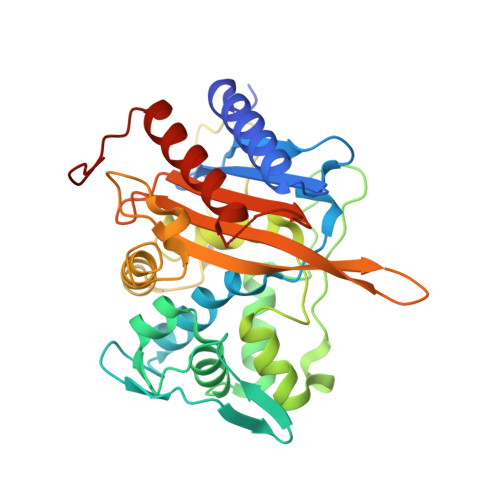
Structural Effect of the Asp345a Insertion in Penicillin-Binding Protein 2 from Penicillin-Resistant Strains of Neisseria gonorrhoeae.
Fedarovich, A., Cook, E., Tomberg, J., Nicholas, R.A., Davies, C.(2014) Biochemistry 53: 7596-7603
- PubMed: 25403720 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi5011317
- Primary Citation Related Structures:
4U3T - PubMed Abstract:
A hallmark of penicillin-binding protein 2 (PBP2) from penicillin-resistant strains of Neisseria gonorrhoeae is insertion of an aspartate after position 345. The insertion resides on a loop near the active site and is immediately adjacent to an existing aspartate (Asp346) that forms a functionally important hydrogen bond with Ser363 of the SxN conserved motif. Insertion of other amino acids, including Glu and Asn, can also lower the rate of acylation by penicillin, but these insertions abolish transpeptidase function. Although the kinetic consequences of the Asp insertion are well-established, how it impacts the structure of PBP2 is unknown. Here, we report the 2.2 Å resolution crystal structure of a truncated construct of PBP2 containing all five mutations present in PBP2 from the penicillin-resistant strain 6140, including the Asp insertion. Commensurate with the strict specificity for the Asp insertion over similar amino acids, the insertion does not cause disordering of the structure, but rather induces localized flexibility in the β2c-β2d loop. The crystal structure resolves the ambiguity of whether the insertion is Asp345a or Asp346a (due to the adjacent Asp) because the hydrogen bond between Asp346 and Ser362 is preserved and the insertion is therefore Asp346a. The side chain of Asp346a projects directly toward the β-lactam-binding site near Asn364 of the SxN motif. The Asp insertion may lower the rate of acylation by sterically impeding binding of the antibiotic or by hindering breakage of the β-lactam ring during acylation because of the negative charge of its side chain.
- Department of Biochemistry and Molecular Biology, Medical University of South Carolina , Charleston, South Carolina 29425, United States.
Organizational Affiliation: