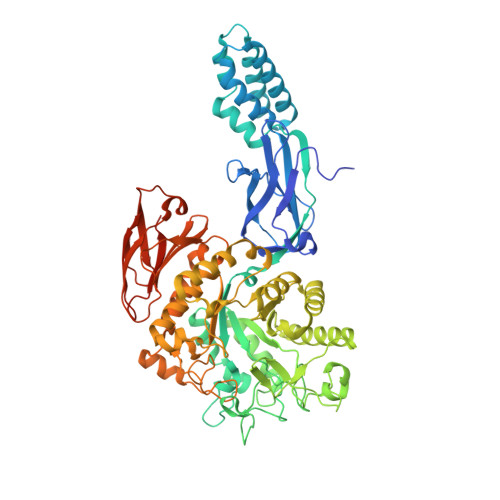
Synthesis of 2-deoxy-2,2-difluoro-alpha-maltosyl fluoride and its X-ray structure in complex with Streptomyces coelicolor GlgEI-V279S.
Thanna, S., Lindenberger, J.J., Gaitonde, V.V., Ronning, D.R., Sucheck, S.J.(2015) Org Biomol Chem 13: 7542-7550
- PubMed: 26072729 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/c5ob00867k
- Primary Citation Related Structures:
4U2Z - PubMed Abstract:
Streptomyces coelicolor (Sco) GlgEI is a glycoside hydrolase involved in α-glucan biosynthesis and can be used as a model enzyme for structure-based inhibitor design targeting Mycobacterium tuberculosis (Mtb) GlgE. The latter is a genetically validated drug target for the development of anti-Tuberculosis (TB) treatments. Inhibition of Mtb GlgE results in a lethal buildup of the GlgE substrate maltose-1-phosphate (M1P). However, Mtb GlgE is difficult to crystallize and affords lower resolution X-ray structures. Sco GlgEI-V279S on the other hand crystallizes readily, produces high resolution X-ray data, and has active site topology identical to Mtb GlgE. We report the X-ray structure of Sco GlgEI-V279S in complex with 2-deoxy-2,2-difluoro-α-maltosyl fluoride (α-MTF, 5) at 2.3 Å resolution. α-MTF was designed as a non-hydrolysable mimic of M1P to probe the active site of GlgE1 prior to covalent bond formation without disruption of catalytic residues. The α-MTF complex revealed hydrogen bonding between Glu423 and the C1F which provides evidence that Glu423 functions as proton donor during catalysis. Further, hydrogen bonding between Arg392 and the axial C2 difluoromethylene moiety of α-MTF was observed suggesting that the C2 position tolerates substitution with hydrogen bond acceptors. The key step in the synthesis of α-MDF was transformation of peracetylated 2-fluoro-maltal 1 into peracetylated 2,2-difluoro-α-maltosyl fluoride 2 in a single step via the use of Selectfluor®.
- Department of Chemistry and Biochemistry, The University of Toledo, 2801 W. Bancroft Street, MS602, Toledo, OH, USA 43606. donald.ronning@utoledo.edu steve.sucheck@utoledo.edu.
Organizational Affiliation: