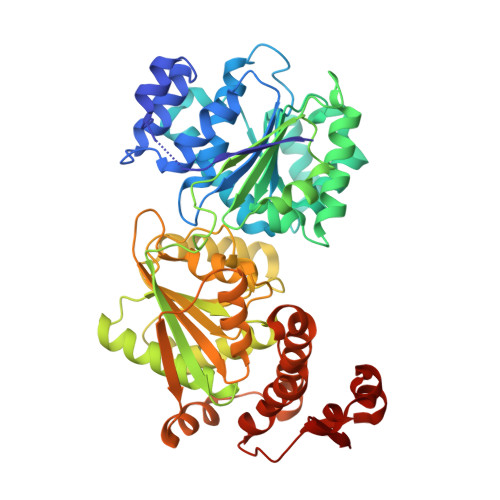
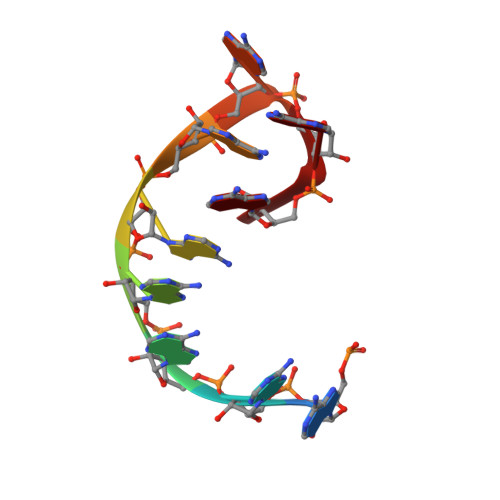
Molecular insights into RNA and DNA helicase evolution from the determinants of specificity for a DEAD-box RNA helicase.
Mallam, A.L., Sidote, D.J., Lambowitz, A.M.(2014) Elife 3: e04630-e04630
- PubMed: 25497230 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.04630
- Primary Citation Related Structures:
4TYN, 4TYW, 4TYY, 4TZ0, 4TZ6 - PubMed Abstract:
How different helicase families with a conserved catalytic 'helicase core' evolved to function on varied RNA and DNA substrates by diverse mechanisms remains unclear. In this study, we used Mss116, a yeast DEAD-box protein that utilizes ATP to locally unwind dsRNA, to investigate helicase specificity and mechanism. Our results define the molecular basis for the substrate specificity of a DEAD-box protein. Additionally, they show that Mss116 has ambiguous substrate-binding properties and interacts with all four NTPs and both RNA and DNA. The efficiency of unwinding correlates with the stability of the 'closed-state' helicase core, a complex with nucleotide and nucleic acid that forms as duplexes are unwound. Crystal structures reveal that core stability is modulated by family-specific interactions that favor certain substrates. This suggests how present-day helicases diversified from an ancestral core with broad specificity by retaining core closure as a common catalytic mechanism while optimizing substrate-binding interactions for different cellular functions.
- Institute for Cellular and Molecular Biology, University of Texas at Austin, Austin, United States.
Organizational Affiliation: