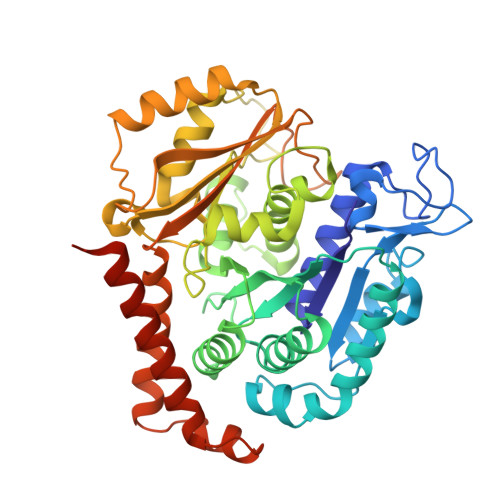
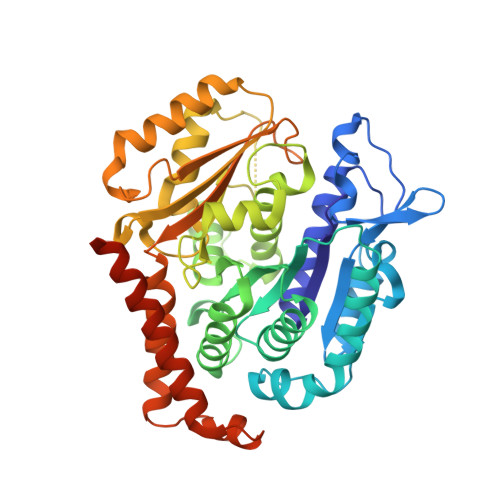
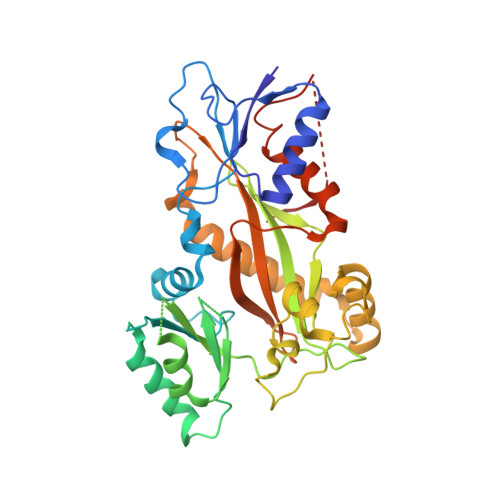
A new tubulin-binding site and pharmacophore for microtubule-destabilizing anticancer drugs.
Prota, A.E., Bargsten, K., Diaz, J.F., Marsh, M., Cuevas, C., Liniger, M., Neuhaus, C., Andreu, J.M., Altmann, K.H., Steinmetz, M.O.(2014) Proc Natl Acad Sci U S A 111: 13817-13821
- PubMed: 25114240 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1408124111
- Primary Citation Related Structures:
4TUY, 4TV8, 4TV9 - PubMed Abstract:
The recent success of antibody-drug conjugates (ADCs) in the treatment of cancer has led to a revived interest in microtubule-destabilizing agents. Here, we determined the high-resolution crystal structure of the complex between tubulin and maytansine, which is part of an ADC that is approved by the US Food and Drug Administration (FDA) for the treatment of advanced breast cancer. We found that the drug binds to a site on β-tubulin that is distinct from the vinca domain and that blocks the formation of longitudinal tubulin interactions in microtubules. We also solved crystal structures of tubulin in complex with both a variant of rhizoxin and the phase 1 drug PM060184. Consistent with biochemical and mutagenesis data, we found that the two compounds bound to the same site as maytansine and that the structures revealed a common pharmacophore for the three ligands. Our results delineate a distinct molecular mechanism of action for the inhibition of microtubule assembly by clinically relevant agents. They further provide a structural basis for the rational design of potent microtubule-destabilizing agents, thus opening opportunities for the development of next-generation ADCs for the treatment of cancer.
- Department of Biology and Chemistry, Laboratory of Biomolecular Research, Paul Scherrer Institut, 5232 Villigen PSI, Switzerland;
Organizational Affiliation: