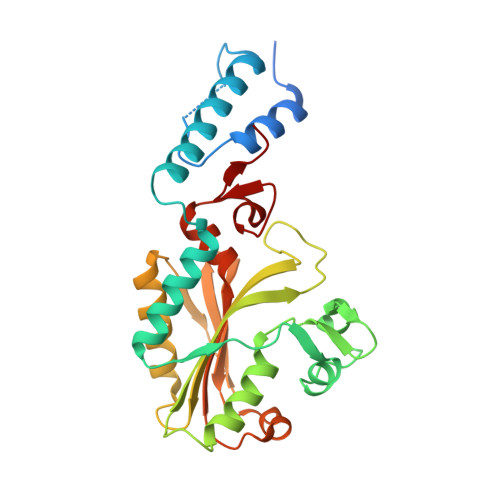
Polyspecific pyrrolysyl-tRNA synthetases from directed evolution.
Guo, L.T., Wang, Y.S., Nakamura, A., Eiler, D., Kavran, J.M., Wong, M., Kiessling, L.L., Steitz, T.A., O'Donoghue, P., Soll, D.(2014) Proc Natl Acad Sci U S A 111: 16724-16729
- PubMed: 25385624 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1419737111
- Primary Citation Related Structures:
4Q6G, 4TQD, 4TQF - PubMed Abstract:
Pyrrolysyl-tRNA synthetase (PylRS) and its cognate tRNA(Pyl) have emerged as ideal translation components for genetic code innovation. Variants of the enzyme facilitate the incorporation >100 noncanonical amino acids (ncAAs) into proteins. PylRS variants were previously selected to acylate N(ε)-acetyl-Lys (AcK) onto tRNA(Pyl). Here, we examine an N(ε)-acetyl-lysyl-tRNA synthetase (AcKRS), which is polyspecific (i.e., active with a broad range of ncAAs) and 30-fold more efficient with Phe derivatives than it is with AcK. Structural and biochemical data reveal the molecular basis of polyspecificity in AcKRS and in a PylRS variant [iodo-phenylalanyl-tRNA synthetase (IFRS)] that displays both enhanced activity and substrate promiscuity over a chemical library of 313 ncAAs. IFRS, a product of directed evolution, has distinct binding modes for different ncAAs. These data indicate that in vivo selections do not produce optimally specific tRNA synthetases and suggest that translation fidelity will become an increasingly dominant factor in expanding the genetic code far beyond 20 amino acids.
- Departments of Molecular Biophysics & Biochemistry and.
Organizational Affiliation: