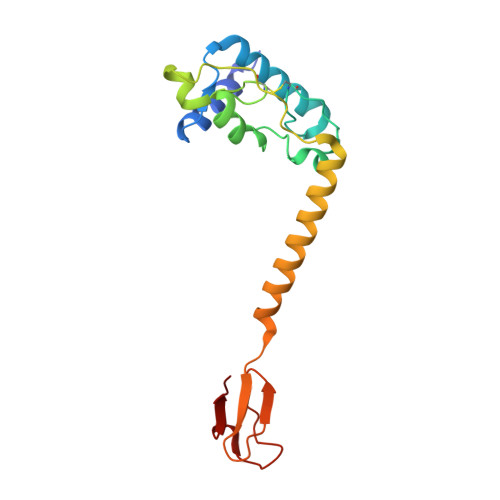
Structural and Biochemical Characterization of a Ferredoxin:Thioredoxin Reductase-like Enzyme from Methanosarcina acetivorans.
Kumar, A.K., Kumar, R.S., Yennawar, N.H., Yennawar, H.P., Ferry, J.G.(2015) Biochemistry 54: 3122-3128
- PubMed: 25915695 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.5b00137
- Primary Citation Related Structures:
4TPU - PubMed Abstract:
Bioinformatics analyses predict the distribution in nature of several classes of diverse disulfide reductases that evolved from an ancestral plant-type ferredoxin:thioredoxin reductase (FTR) catalytic subunit to meet a variety of ecological needs. Methanosarcina acetivorans is a methane-producing species from the domain Archaea predicted to encode an FTR-like enzyme with two domains, one resembling the FTR catalytic subunit and the other containing a rubredoxin-like domain replacing the variable subunit of present-day FTR enzymes. M. acetivorans is of special interest as it was recently proposed to have evolved at the time of the end-Permian extinction and to be largely responsible for the most severe biotic crisis in the fossil record by converting acetate to methane. The crystal structure and biochemical characteristics were determined for the FTR-like enzyme from M. acetivorans, here named FDR (ferredoxin disulfide reductase). The results support a role for the rubredoxin-like center of FDR in transfer of electrons from ferredoxin to the active-site [Fe₄S₄] cluster adjacent to a pair of redox-active cysteines participating in reduction of disulfide substrates. A mechanism is proposed for disulfide reduction similar to one of two mechanisms previously proposed for the plant-type FTR. Overall, the results advance the biochemical and evolutionary understanding of the FTR-like family of enzymes and the conversion of acetate to methane that is an essential link in the global carbon cycle and presently accounts for most of this greenhouse gas that is biologically generated.