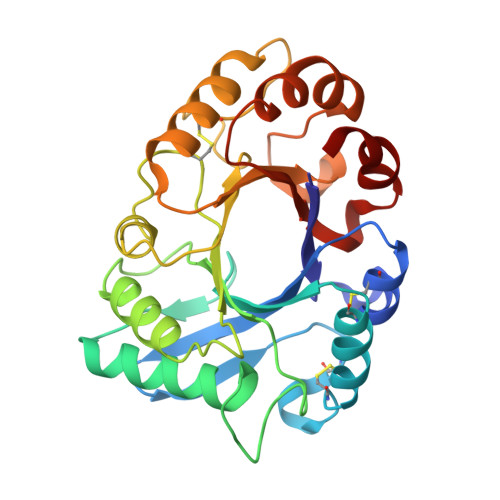
Crystal structure of class III chitinase from pomegranate provides the insight into its metal storage capacity.
Masuda, T., Zhao, G., Mikami, B.(2015) Biosci Biotechnol Biochem 79: 45-50
- PubMed: 25252615 Search on PubMed
- DOI: https://doi.org/10.1080/09168451.2014.962475
- Primary Citation Related Structures:
4TOQ - PubMed Abstract:
Chitinase hydrolyzes the β-1,4-glycosidic bond in chitin. In higher plants, this enzyme has been regarded as a pathogenesis-related protein. Recently, we identified a class III chitinase, which functions as a calcium storage protein in pomegranate (Punica granatum) seed (PSC, pomegranate seed chitinase). Here, we solved a crystal structure of PSC at 1.6 Å resolution. Although its overall structure, including the structure of catalytic site and non-proline cis-peptides, was closely similar to those of other class III chitinases, PSC had some unique structural characteristics. First, there were some metal-binding sites with coordinated water molecules on the surface of PSC. Second, many unconserved aspartate residues were present in the PSC sequence which rendered the surface of PSC negatively charged. This acidic electrostatic property is in contrast to that of hevamine, well-characterized plant class III chitinase, which has rather a positively charged surface. Thus, the crystal structure provides a clue for metal association property of PSC.
- a Laboratory of Food Quality Design and Development, Division of Agronomy and Horticultural Science, Graduate School of Agriculture , Kyoto University , Gokasho, Uji, Kyoto , Japan.
Organizational Affiliation: