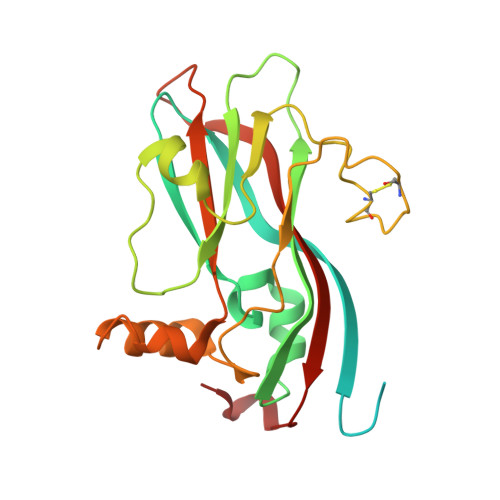
Refined structure of southern bean mosaic virus at 2.9 A resolution.
Silva, A.M., Rossmann, M.G.(1987) J Mol Biology 197: 69-87
- PubMed: 3681993 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(87)90610-3
- Primary Citation Related Structures:
4SBV - PubMed Abstract:
The T = 3 capsid of southern bean mosaic virus is analyzed in detail. The beta-sheets of the beta-barrel folding motif that form the subunits show a high degree of twist, generated by several beta-bulges. Only 34 water molecules were identified in association with the three quasi-equivalent subunits, most of them on the external viral surface. Subunit contacts related by quasi-3-fold axes are similar, are dominated by polar interactions and have almost identical calcium binding sites. There is no metal ion on the quasi-3-fold axis, as previously reported. Subunits related by quasi-2-fold and icosahedral 2-fold axes have different contacts but nevertheless display almost identical interactions between the antiparallel helices alpha A. A dipole-dipole type interaction between these helices may produce an energetically stable hinge that allows two types of dimers in a T = 3 assembly. The temperature factor distribution, the hydrogen-bonding pattern, and the contacts across the icosahedral 2-fold axes suggest that one of the dimer types is present in the intact virion and probably also in solution; the other is produced only during capsid assembly. Interactions along the 5-fold axes are mainly polar and possibly form an ion channel. The beta-sheet structures of the three subunits can be superimposed with considerable precision. Significant relative distortions between quasi-equivalent subunits occur mainly in helices and loops. The two dimeric forms and the subunit distortions are the consequence of the non-equivalent subunit environments in the capsid.
- Departamento de Fisica, Facultad de Ciencias Exactas, Universidad Nacional de La Plata, Argentina.
Organizational Affiliation: