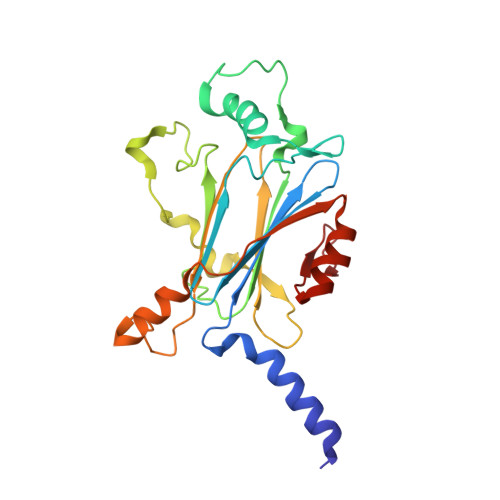
Structure of CPV17 polyhedrin determined by the improved analysis of serial femtosecond crystallographic data.
Ginn, H.M., Messerschmidt, M., Ji, X., Zhang, H., Axford, D., Gildea, R.J., Winter, G., Brewster, A.S., Hattne, J., Wagner, A., Grimes, J.M., Evans, G., Sauter, N.K., Sutton, G., Stuart, D.I.(2015) Nat Commun 6: 6435-6435
- PubMed: 25751308 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms7435
- Primary Citation Related Structures:
4S1K, 4S1L - PubMed Abstract:
The X-ray free-electron laser (XFEL) allows the analysis of small weakly diffracting protein crystals, but has required very many crystals to obtain good data. Here we use an XFEL to determine the room temperature atomic structure for the smallest cytoplasmic polyhedrosis virus polyhedra yet characterized, which we failed to solve at a synchrotron. These protein microcrystals, roughly a micron across, accrue within infected cells. We use a new physical model for XFEL diffraction, which better estimates the experimental signal, delivering a high-resolution XFEL structure (1.75 Å), using fewer crystals than previously required for this resolution. The crystal lattice and protein core are conserved compared with a polyhedrin with less than 10% sequence identity. We explain how the conserved biological phenotype, the crystal lattice, is maintained in the face of extreme environmental challenge and massive evolutionary divergence. Our improved methods should open up more challenging biological samples to XFEL analysis.
- Division of Structural Biology, The Wellcome Trust Centre for Human Genetics, University of Oxford, Roosevelt Drive, Oxford, Oxfordshire OX3 7BN, UK.
Organizational Affiliation: