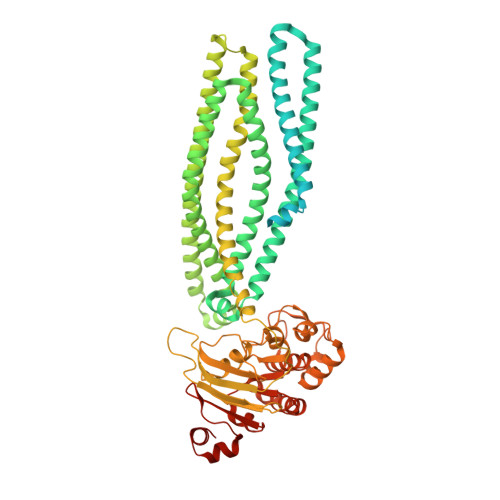
Crystal structures of a polypeptide processing and secretion transporter.
Lin, D.Y., Huang, S., Chen, J.(2015) Nature 523: 425-430
- PubMed: 26201595 Search on PubMed
- DOI: https://doi.org/10.1038/nature14623
- Primary Citation Related Structures:
4RY2, 4S0F - PubMed Abstract:
Bacteria secrete peptides and proteins to communicate, to poison competitors, and to manipulate host cells. Among the various protein-translocation machineries, the peptidase-containing ATP-binding cassette transporters (PCATs) are appealingly simple. Each PCAT contains two peptidase domains that cleave the secretion signal from the substrate, two transmembrane domains that form a translocation pathway, and two nucleotide-binding domains that hydrolyse ATP. In Gram-positive bacteria, PCATs function both as maturation proteases and exporters for quorum-sensing or antimicrobial polypeptides. In Gram-negative bacteria, PCATs interact with two other membrane proteins to form the type 1 secretion system. Here we present crystal structures of PCAT1 from Clostridium thermocellum in two different conformations. These structures, accompanied by biochemical data, show that the translocation pathway is a large α-helical barrel sufficient to accommodate small folded proteins. ATP binding alternates access to the transmembrane pathway and also regulates the protease activity, thereby coupling substrate processing to translocation.
- 1] Laboratory of Membrane Biology and Biophysics, The Rockefeller University, 1230 York Avenue, New York, New York 10065, USA [2] Howard Hughes Medical Institute, 1230 York Avenue, New York, New York 10065, USA.
Organizational Affiliation: