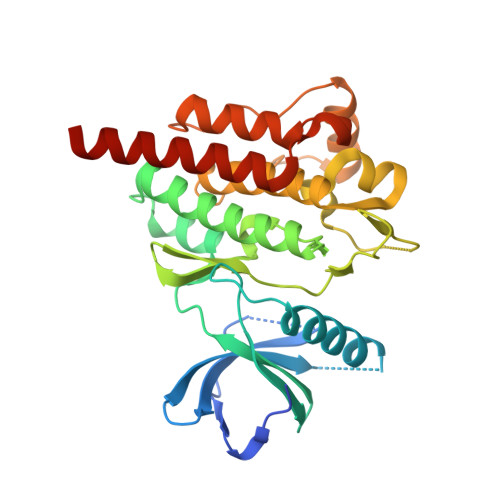
Discovery and profiling of a selective and efficacious syk inhibitor.
Thoma, G., Smith, A.B., van Eis, M.J., Vangrevelinghe, E., Blanz, J., Aichholz, R., Littlewood-Evans, A., Lee, C.C., Liu, H., Zerwes, H.G.(2015) J Med Chem 58: 1950-1963
- PubMed: 25633741 Search on PubMed
- DOI: https://doi.org/10.1021/jm5018863
- Primary Citation Related Structures:
4RX7, 4RX8, 4RX9 - PubMed Abstract:
We describe the discovery of selective and potent Syk inhibitor 11, which exhibited favorable PK profiles in rat and dog and was found to be active in a collagen-induced arthritis model in rats. Compound 11 was selected for further profiling, but, unfortunately, in GLP toxicological studies it showed liver findings in rat and dog. Nevertheless, 11 could become a valuable tool compound to investigate the rich biology of Syk in vitro and in vivo.
- Global Discovery Chemistry, ‡Analytical Sciences & Imaging, §Autoimmunity, Transplantation and Inflammation Research, Novartis Institutes for Biomedical Research , 4056 Basel, Switzerland.
Organizational Affiliation: