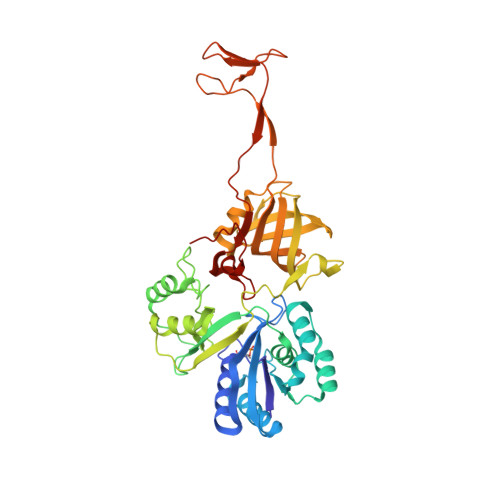
A covalent approach for site-specific RNA labeling in Mammalian cells.
Li, F., Dong, J., Hu, X., Gong, W., Li, J., Shen, J., Tian, H., Wang, J.(2015) Angew Chem Int Ed Engl 54: 4597-4602
- PubMed: 25694369 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201410433
- Primary Citation Related Structures:
4RVZ - PubMed Abstract:
Advances in RNA research and RNA nanotechnology depend on the ability to manipulate and probe RNA with high precision through chemical approaches, both in vitro and in mammalian cells. However, covalent RNA labeling methods with scope and versatility comparable to those of current protein labeling strategies are underdeveloped. A method is reported for the site- and sequence-specific covalent labeling of RNAs in mammalian cells by using tRNA(Ile2) -agmatidine synthetase (Tias) and click chemistry. The crystal structure of Tias in complex with an azide-bearing agmatine analogue was solved to unravel the structural basis for Tias/substrate recognition. The unique RNA sequence specificity and plastic Tias/substrate recognition enable the site-specific transfer of azide/alkyne groups to an RNA molecule of interest in vitro and in mammalian cells. Subsequent click chemistry reactions facilitate the versatile labeling, functionalization, and visualization of target RNA.
- Laboratory of RNA Biology and Laboratory of Quantum Biophysics, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Chaoyang District, Beijing, 100101 (China).
Organizational Affiliation: