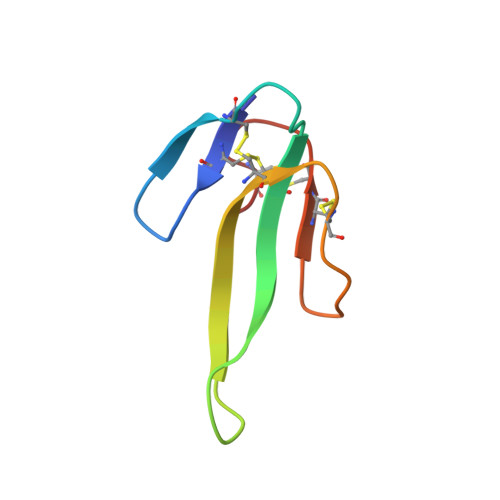
Fulditoxin, representing a new class of dimeric snake toxins, defines novel pharmacology at nicotinic ACh receptors.
Foo, C.S., Jobichen, C., Hassan-Puttaswamy, V., Dekan, Z., Tae, H.S., Bertrand, D., Adams, D.J., Alewood, P.F., Sivaraman, J., Nirthanan, S., Kini, R.M.(2020) Br J Pharmacol 177: 1822-1840
- PubMed: 31877243 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/bph.14954
- Primary Citation Related Structures:
4RUD - PubMed Abstract:
Animal toxins have contributed significantly to our understanding of the neurobiology of receptors and ion channels. We studied the venom of the coral snake Micrurus fulvius fulvius and identified and characterized the structure and pharmacology of a new homodimeric neurotoxin, fulditoxin, that exhibited novel pharmacology at nicotinic ACh receptors (nAChRs). Fulditoxin was isolated by chromatography, chemically synthesized, its structure determined by X-ray crystallography, and its pharmacological actions on nAChRs characterized by organ bath assays and two-electrode voltage clamp electrophysiology. Fulditoxin's distinct 1.95-Å quaternary structure revealed two short-chain three-finger α-neurotoxins (α-3FNTxs) non-covalently bound by hydrophobic interactions and an ability to bind metal and form tetrameric complexes, not reported previously for three-finger proteins. Although fulditoxin lacked all conserved amino acids canonically important for inhibiting nAChRs, it produced postsynaptic neuromuscular blockade of chick muscle at nanomolar concentrations, comparable to the prototypical α-bungarotoxin. This neuromuscular blockade was completely reversible, which is unusual for snake α-3FNTxs. Fulditoxin, therefore, interacts with nAChRs by utilizing a different pharmacophore. Unlike short-chain α-3FNTxs that bind only to muscle nAChRs, fulditoxin utilizes dimerization to expand its pharmacological targets to include human neuronal α4β2, α7, and α3β2 nAChRs which it blocked with IC 50 values of 1.8, 7, and 12 μM respectively. Based on its distinct quaternary structure and unusual pharmacology, we named this new class of dimeric Micrurus neurotoxins represented by fulditoxin as Σ-neurotoxins, which offers greater insight into understanding the interactions between nAChRs and peptide antagonists.
- Protein Science Laboratory, Department of Biological Sciences, Faculty of Science, National University of Singapore, Singapore, Singapore.
Organizational Affiliation: