8-Tetrahydropyran-2-yl Chromans: Highly Selective Beta-Site Amyloid Precursor Protein Cleaving Enzyme 1 (BACE1) Inhibitors.
Thomas, A.A., Hunt, K.W., Newhouse, B., Watts, R.J., Liu, X., Vigers, G., Smith, D., Rhodes, S.P., Brown, K.D., Otten, J.N., Burkard, M., Cox, A.A., Geck Do, M.K., Dutcher, D., Rana, S., DeLisle, R.K., Regal, K., Wright, A.D., Groneberg, R., Liao, J., Scearce-Levie, K., Siu, M., Purkey, H.E., Lyssikatos, J.P.(2014) J Med Chem 57: 10112-10129
- PubMed: 25411915
- DOI: https://doi.org/10.1021/jm5015132
- Primary Citation of Related Structures:
4R5N, 4RRN, 4RRO, 4RRS - PubMed Abstract:
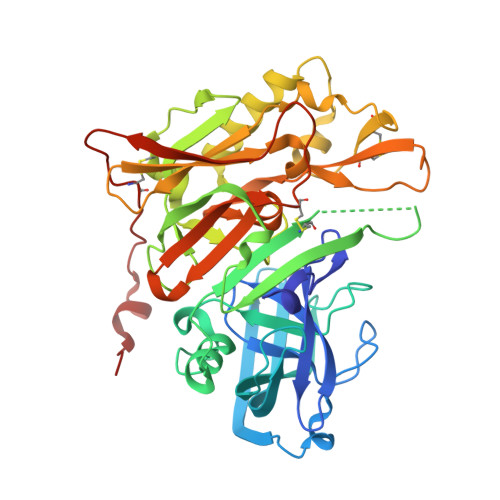
A series of 2,3,4,4a,10,10a-hexahydropyrano[3,2-b]chromene analogs was developed that demonstrated high selectivity (>2000-fold) for BACE1 vs Cathepsin D (CatD). Three different Asp-binding moieties were examined: spirocyclic acyl guanidines, aminooxazolines, and aminothiazolines in order to modulate potency, selectivity, efflux, and permeability. Guided by structure based design, changes to P2' and P3 moieties were explored. A conformationally restricted P2' methyl group provided inhibitors with excellent cell potency (37-137 nM) and selectivity (435 to >2000-fold) for BACE1 vs CatD. These efforts lead to compound 59, which demonstrated a 69% reduction in rat CSF Aβ1-40 at 60 mg/kg (PO).
- Array BioPharma, 3200 Walnut Street, Boulder, Colorado 80301, United States.
Organizational Affiliation: