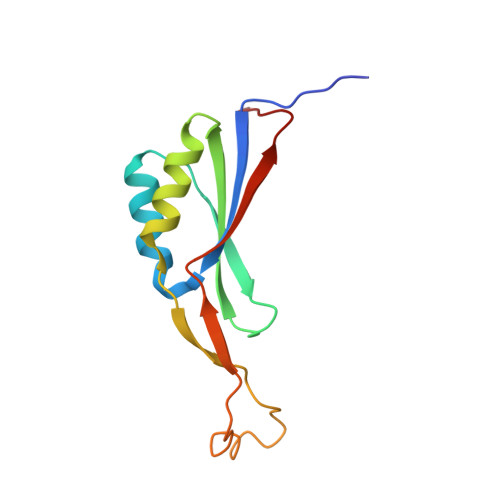
Identification, Characterization, and Structure Analysis of the Cyclic di-AMP-binding PII-like Signal Transduction Protein DarA.
Gundlach, J., Dickmanns, A., Schroder-Tittmann, K., Neumann, P., Kaesler, J., Kampf, J., Herzberg, C., Hammer, E., Schwede, F., Kaever, V., Tittmann, K., Stulke, J., Ficner, R.(2015) J Biological Chem 290: 3069-3080
- PubMed: 25433025 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.619619
- Primary Citation Related Structures:
4RLE - PubMed Abstract:
The cyclic dimeric AMP nucleotide c-di-AMP is an essential second messenger in Bacillus subtilis. We have identified the protein DarA as one of the prominent c-di-AMP receptors in B. subtilis. Crystal structure analysis shows that DarA is highly homologous to PII signal transducer proteins. In contrast to PII proteins, the functionally important B- and T-loops are swapped with respect to their size. DarA is a homotrimer that binds three molecules of c-di-AMP, each in a pocket located between two subunits. We demonstrate that DarA is capable to bind c-di-AMP and with lower affinity cyclic GMP-AMP (3'3'-cGAMP) but not c-di-GMP or 2'3'-cGAMP. Consistently the crystal structure shows that within the ligand-binding pocket only one adenine is highly specifically recognized, whereas the pocket for the other adenine appears to be promiscuous. Comparison with a homologous ligand-free DarA structure reveals that c-di-AMP binding is accompanied by conformational changes of both the fold and the position of the B-loop in DarA.
- From the Departments of General Microbiology.
Organizational Affiliation: