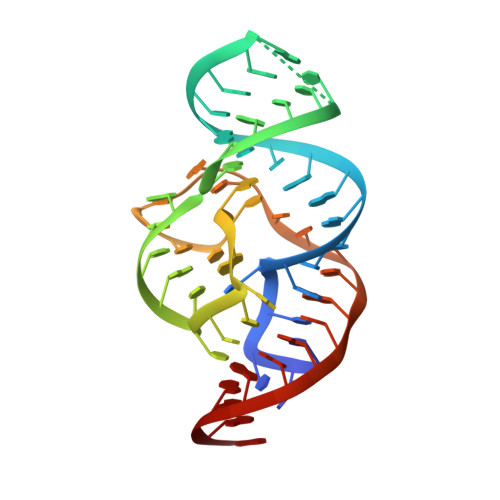
In-line alignment and Mg(2+) coordination at the cleavage site of the env22 twister ribozyme.
Ren, A., Kosutic, M., Rajashankar, K.R., Frener, M., Santner, T., Westhof, E., Micura, R., Patel, D.J.(2014) Nat Commun 5: 5534-5534
- PubMed: 25410397 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms6534
- Primary Citation Related Structures:
4RGE, 4RGF - PubMed Abstract:
Small self-cleaving nucleolytic ribozymes contain catalytic domains that accelerate site-specific cleavage/ligation of phosphodiester backbones. We report on the 2.9-Å crystal structure of the env22 twister ribozyme, which adopts a compact tertiary fold stabilized by co-helical stacking, double-pseudoknot formation and long-range pairing interactions. The U-A cleavage site adopts a splayed-apart conformation with the modelled 2'-O of U positioned for in-line attack on the adjacent to-be-cleaved P-O5' bond. Both an invariant guanosine and a Mg(2+) are directly coordinated to the non-bridging phosphate oxygens at the U-A cleavage step, with the former positioned to contribute to catalysis and the latter to structural integrity. The impact of key mutations on cleavage activity identified an invariant guanosine that contributes to catalysis. Our structure of the in-line aligned env22 twister ribozyme is compared with two recently reported twister ribozymes structures, which adopt similar global folds, but differ in conformational features around the cleavage site.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, New York 10065, USA.
Organizational Affiliation: