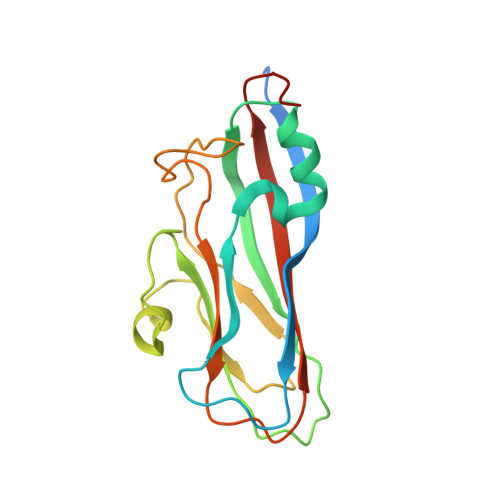
Crystal Structures of a Piscine Betanodavirus: Mechanisms of Capsid Assembly and Viral Infection
Chen, N.C., Yoshimura, M., Guan, H.H., Wang, T.Y., Misumi, Y., Lin, C.C., Chuankhayan, P., Nakagawa, A., Chan, S.I., Tsukihara, T., Chen, T.Y., Chen, C.J.(2015) PLoS Pathog 11: e1005203-e1005203
- PubMed: 26491970 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1005203
- Primary Citation Related Structures:
4RFT, 4RFU, 4WIZ - PubMed Abstract:
Betanodaviruses cause massive mortality in marine fish species with viral nervous necrosis. The structure of a T = 3 Grouper nervous necrosis virus-like particle (GNNV-LP) is determined by the ab initio method with non-crystallographic symmetry averaging at 3.6 Å resolution. Each capsid protein (CP) shows three major domains: (i) the N-terminal arm, an inter-subunit extension at the inner surface; (ii) the shell domain (S-domain), a jelly-roll structure; and (iii) the protrusion domain (P-domain) formed by three-fold trimeric protrusions. In addition, we have determined structures of the T = 1 subviral particles (SVPs) of (i) the delta-P-domain mutant (residues 35-217) at 3.1 Å resolution; and (ii) the N-ARM deletion mutant (residues 35-338) at 7 Å resolution; and (iii) the structure of the individual P-domain (residues 214-338) at 1.2 Å resolution. The P-domain reveals a novel DxD motif asymmetrically coordinating two Ca2+ ions, and seems to play a prominent role in the calcium-mediated trimerization of the GNNV CPs during the initial capsid assembly process. The flexible N-ARM (N-terminal arginine-rich motif) appears to serve as a molecular switch for T = 1 or T = 3 assembly. Finally, we find that polyethylene glycol, which is incorporated into the P-domain during the crystallization process, enhances GNNV infection. The present structural studies together with the biological assays enhance our understanding of the role of the P-domain of GNNV in the capsid assembly and viral infection by this betanodavirus.
- Institute of Biotechnology, National Cheng Kung University, Tainan, Taiwan; Life Science Group, Scientific Research Division, National Synchrotron Radiation Research Center, Hsinchu, Taiwan.
Organizational Affiliation: