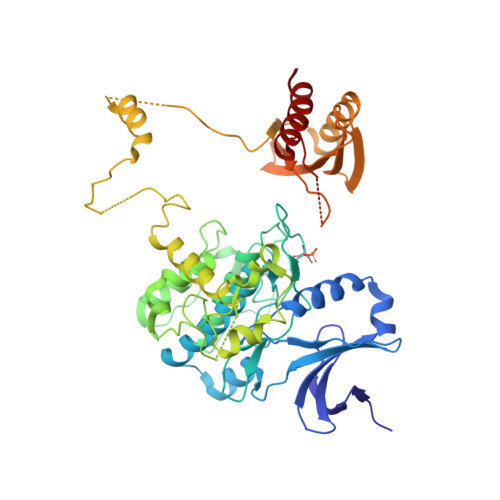
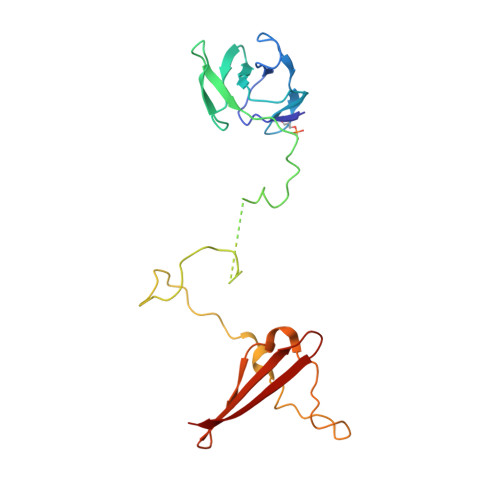
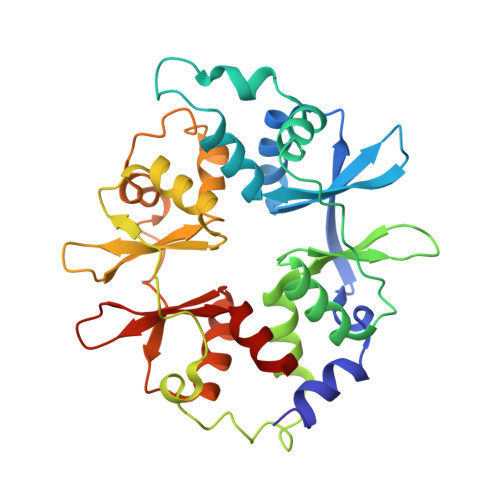
Structural basis of AMPK regulation by adenine nucleotides and glycogen.
Li, X., Wang, L., Zhou, X.E., Ke, J., de Waal, P.W., Gu, X., Tan, M.H., Wang, D., Wu, D., Xu, H.E., Melcher, K.(2015) Cell Res 25: 50-66
- PubMed: 25412657 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/cr.2014.150
- Primary Citation Related Structures:
4RED, 4RER, 4REW - PubMed Abstract:
AMP-activated protein kinase (AMPK) is a central cellular energy sensor and regulator of energy homeostasis, and a promising drug target for the treatment of diabetes, obesity, and cancer. Here we present low-resolution crystal structures of the human α1β2γ1 holo-AMPK complex bound to its allosteric modulators AMP and the glycogen-mimic cyclodextrin, both in the phosphorylated (4.05 Å) and non-phosphorylated (4.60 Å) state. In addition, we have solved a 2.95 Å structure of the human kinase domain (KD) bound to the adjacent autoinhibitory domain (AID) and have performed extensive biochemical and mutational studies. Together, these studies illustrate an underlying mechanism of allosteric AMPK modulation by AMP and glycogen, whose binding changes the equilibria between alternate AID (AMP) and carbohydrate-binding module (glycogen) interactions.
- 1] Key Laboratory of Regenerative Biology, Guangzhou Institute of Biomedicine and Health, Chinese Academy of Sciences, Guangzhou, Guangdong 510530, China [2] School of Life Science, University of Science and Technology of China, Hefei, Anhui 230027, China [3] Laboratory of Structural Sciences, Van Andel Research Institute, 333 Bostwick Ave, NE, Grand Rapids, MI 49503, USA.
Organizational Affiliation: