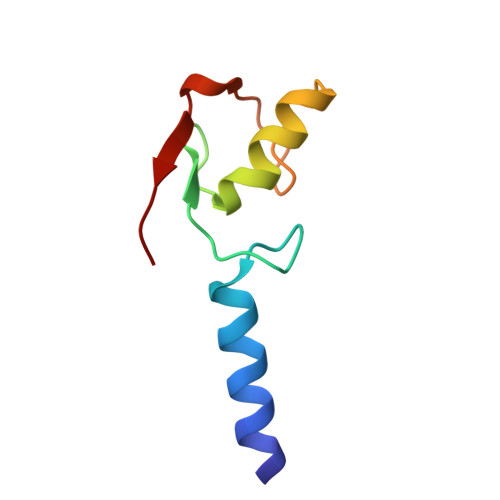
Structure of the yeast Bre1 RING domain.
Kumar, P., Wolberger, C.(2015) Proteins 83: 1185-1190
- PubMed: 25864391 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.24812
- Primary Citation Related Structures:
4R7E - PubMed Abstract:
Monoubiquitination of histone H2B at Lys123 in yeast plays a critical role in regulating transcription, mRNA export, DNA replication, and the DNA damage response. The RING E3 ligase, Bre1, catalyzes monoubiquitination of H2B in concert with the E2 ubiquitin-conjugating enzyme, Rad6. The crystal structure of a C-terminal fragment of Bre1 shows that the catalytic RING domain is preceded by an N-terminal helix that mediates coiled-coil interactions with a crystallographically related monomer. Homology modeling suggests that the human homologue of Bre1, RNF20/RNF40, heterodimerizes through similar coiled-coil interactions.
- Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, 725 North Wolfe Street, Baltimore, Maryland, 21205.
Organizational Affiliation: