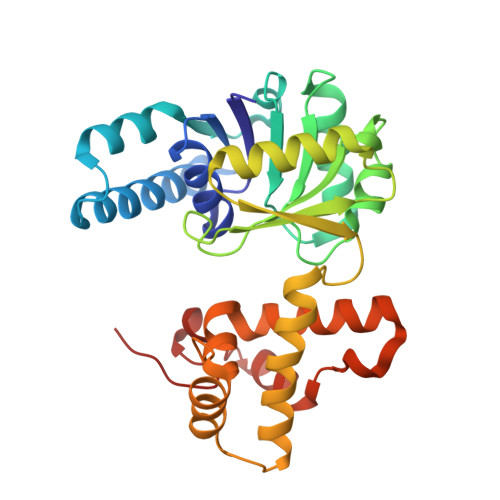
Crystal structure of (S)-3-hydroxybutyryl-CoA dehydrogenase from Clostridium butyricum and its mutations that enhance reaction kinetics
Kim, E.J., Kim, J., Ahn, J.W., Kim, Y.J., Chang, J.H., Kim, K.J.(2014) J Microbiol Biotechnol 24: 1636-1643
- PubMed: 25112316 Search on PubMed
- DOI: https://doi.org/10.4014/jmb.1407.07027
- Primary Citation Related Structures:
4R1N - PubMed Abstract:
3-Hydroxybutyryl-CoA dehydrogenase is an enzyme that catalyzes the second step in the biosynthesis of n-butanol from acetyl-CoA, in which acetoacetyl-CoA is reduced to 3-hydroxybutyryl-CoA. To understand the molecular mechanisms of n-butanol biosynthesis, we determined the crystal structure of 3-hydroxybutyryl-CoA dehydrogenase from Clostridium butyricum (CbHBD). The monomer structure of CbHBD exhibits a two-domain topology, with N- and C-terminal domains, and the dimerization of the enzyme was mostly constituted at the C-terminal domain. The mode of cofactor binding to CbHBD was elucidated by determining the crystal structure of the enzyme in complex with NAD(+). We also determined the enzyme's structure in complex with its acetoacetyl-CoA substrate, revealing that the adenosine diphosphate moiety was not highly stabilized compared with the remainder of the acetoacetyl-CoA molecule. Using this structural information, we performed a series of sitedirected mutagenesis experiments on the enzyme, such as changing residues located near the substrate-binding site, and finally developed a highly efficient CbHBD K50A/K54A/L232Y triple mutant enzyme that exhibited approximately 5-fold higher enzyme activity than did the wild type. The increased enzyme activity of the mutant was confirmed by enzyme kinetic measurements. The highly efficient mutant enzyme should be useful for increasing the production rate of n-butanol.
- School of Life Sciences, KNU Creative BioResearch Group, Kyungpook National University, Daegu 702-701, Republic of Korea.
Organizational Affiliation: