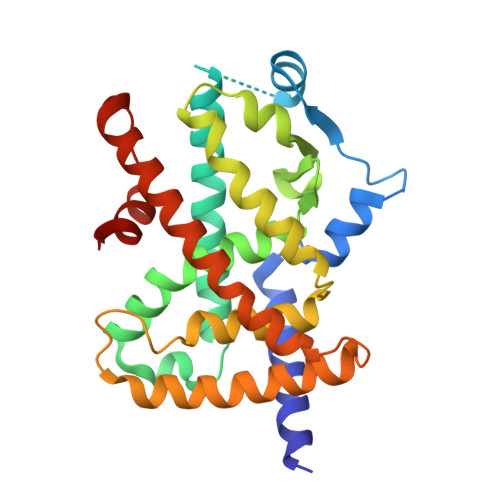
SR2067 Reveals a Unique Kinetic and Structural Signature for PPAR gamma Partial Agonism.
van Marrewijk, L.M., Polyak, S.W., Hijnen, M., Kuruvilla, D., Chang, M.R., Shin, Y., Kamenecka, T.M., Griffin, P.R., Bruning, J.B.(2016) ACS Chem Biol 11: 273-283
- PubMed: 26579553 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschembio.5b00580
- Primary Citation Related Structures:
4R06 - PubMed Abstract:
Synthetic full agonists of PPARγ have been prescribed for the treatment of diabetes due to their ability to regulate glucose homeostasis and insulin sensitization. While the use of full agonists of PPARγ has been hampered due to severe side effects, partial agonists have shown promise due to their decreased incidence of such side effects in preclinical models. No kinetic information has been forthcoming in regard to the mechanism of full versus partial agonism of PPARγ to date. Here, we describe the discovery of a partial agonist, SR2067. A co-crystal structure obtained at 2.2 Å resolution demonstrates that interactions with the β-sheet are driven exclusively via hydrophobic interactions mediated through a naphthalene group, an observation that is unique from other partial agonists. Surface plasmon resonance revealed that SR2067 binds to the receptor with higher affinity (KD = 513 nM) as compared to that of full agonist rosiglitazone, yet it has a much slower off rate compared to that of rosiglitazone.
- School of Biological Sciences, The University of Adelaide , Adelaide, South Australia 5005, Australia.
Organizational Affiliation: