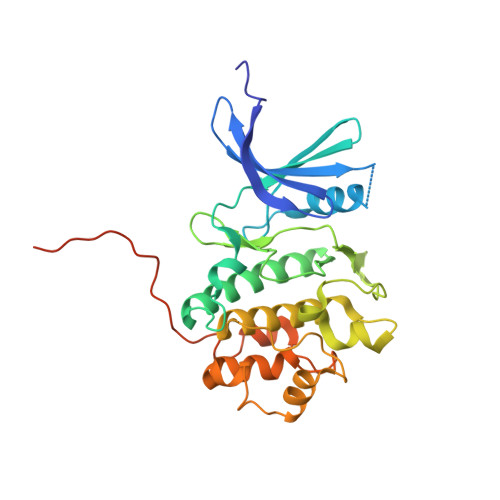
Discovery of the 1,7-diazacarbazole class of inhibitors of checkpoint kinase 1.
Gazzard, L., Appleton, B., Chapman, K., Chen, H., Clark, K., Drobnick, J., Goodacre, S., Halladay, J., Lyssikatos, J., Schmidt, S., Sideris, S., Wiesmann, C., Williams, K., Wu, P., Yen, I., Malek, S.(2014) Bioorg Med Chem Lett 24: 5704-5709
- PubMed: 25453805 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2014.10.063
- Primary Citation Related Structures:
4QYE, 4QYF, 4QYG, 4QYH - PubMed Abstract:
Checkpoint kinase 1 (ChK1) is activated in response to DNA damage, acting to temporarily block cell cycle progression and allow for DNA repair. It is envisaged that inhibition of ChK1 will sensitize tumor cells to treatment with DNA-damaging therapies, and may enhance the therapeutic window. High throughput screening identified carboxylate-containing diarylpyrazines as a prominent hit series, but with limited biochemical potency and no cellular activity. Through a series of SAR investigations and X-ray crystallographic analysis the critical role of polar contacts with conserved waters in the kinase back pocket was established. Structure-based design, guided by in silico modeling, transformed the series to better satisfy these contacts and the novel 1,7-diazacarbazole class of inhibitors was discovered. Here we present the genesis of this novel series and the identification of GNE-783, a potent, selective and orally bioavailable inhibitor of ChK1.
- Department of Discovery Chemistry, Genentech Inc., 1 DNA Way, South San Francisco, CA 94080, United States.
Organizational Affiliation: