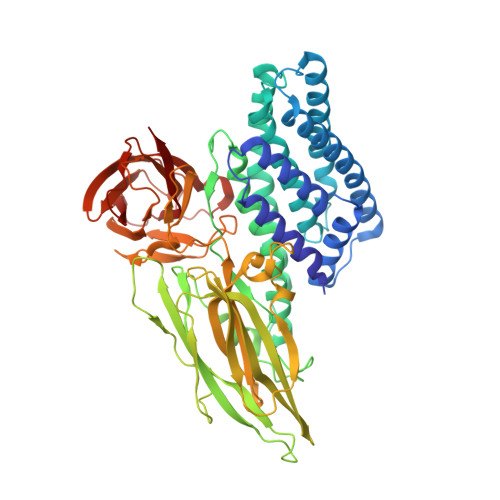
Protein crystal structure obtained at 2.9 angstrom resolution from injecting bacterial cells into an X-ray free-electron laser beam.
Sawaya, M.R., Cascio, D., Gingery, M., Rodriguez, J., Goldschmidt, L., Colletier, J.P., Messerschmidt, M.M., Boutet, S., Koglin, J.E., Williams, G.J., Brewster, A.S., Nass, K., Hattne, J., Botha, S., Doak, R.B., Shoeman, R.L., DePonte, D.P., Park, H.W., Federici, B.A., Sauter, N.K., Schlichting, I., Eisenberg, D.S.(2014) Proc Natl Acad Sci U S A 111: 12769-12774
- PubMed: 25136092 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1413456111
- Primary Citation Related Structures:
4QX0, 4QX1, 4QX2, 4QX3 - PubMed Abstract:
It has long been known that toxins produced by Bacillus thuringiensis (Bt) are stored in the bacterial cells in crystalline form. Here we describe the structure determination of the Cry3A toxin found naturally crystallized within Bt cells. When whole Bt cells were streamed into an X-ray free-electron laser beam we found that scattering from other cell components did not obscure diffraction from the crystals. The resolution limits of the best diffraction images collected from cells were the same as from isolated crystals. The integrity of the cells at the moment of diffraction is unclear; however, given the short time (∼ 5 µs) between exiting the injector to intersecting with the X-ray beam, our result is a 2.9-Å-resolution structure of a crystalline protein as it exists in a living cell. The study suggests that authentic in vivo diffraction studies can produce atomic-level structural information.
- UCLA-DOE Institute for Genomics and Proteomics, Department of Biological Chemistry, and.
Organizational Affiliation: