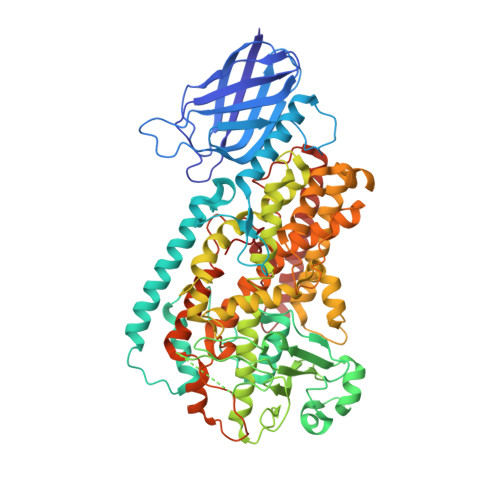
Crystal Structure of a Lipoxygenase in Complex with Substrate: THE ARACHIDONIC ACID-BINDING SITE OF 8R-LIPOXYGENASE.
Neau, D.B., Bender, G., Boeglin, W.E., Bartlett, S.G., Brash, A.R., Newcomer, M.E.(2014) J Biological Chem 289: 31905-31913
- PubMed: 25231982 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.599662
- Primary Citation Related Structures:
4QWT - PubMed Abstract:
Lipoxygenases (LOX) play critical roles in mammalian biology in the generation of potent lipid mediators of the inflammatory response; consequently, they are targets for the development of isoform-specific inhibitors. The regio- and stereo-specificity of the oxygenation of polyunsaturated fatty acids by the enzymes is understood in terms of the chemistry, but structural observation of the enzyme-substrate interactions is lacking. Although several LOX crystal structures are available, heretofore the rapid oxygenation of bound substrate has precluded capture of the enzyme-substrate complex, leaving a gap between chemical and structural insights. In this report, we describe the 2.0 Å resolution structure of 8R-LOX in complex with arachidonic acid obtained under anaerobic conditions. Subtle rearrangements, primarily in the side chains of three amino acids, allow binding of arachidonic acid in a catalytically competent conformation. Accompanying experimental work supports a model in which both substrate tethering and cavity depth contribute to positioning the appropriate carbon at the catalytic machinery.
- Department of Chemistry and Chemical Biology, Cornell University, Northeastern Collaborative Access Team, Argonne National Laboratory, Argonne, Illinois 60439, and.
Organizational Affiliation: