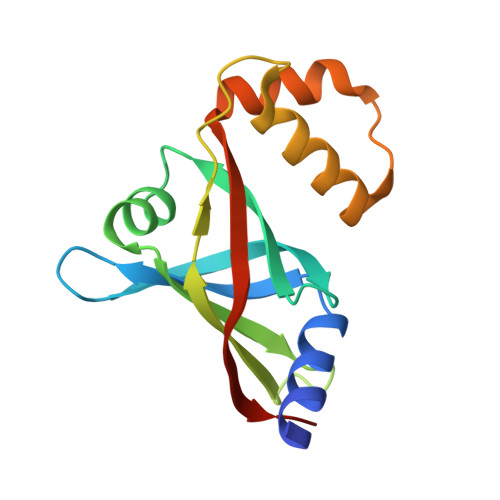
Molecular insights into the binding of coenzyme F420 to the conserved protein Rv1155 from Mycobacterium tuberculosis.
Mashalidis, E.H., Gittis, A.G., Tomczak, A., Abell, C., Barry, C.E., Garboczi, D.N.(2015) Protein Sci 24: 729-740
- PubMed: 25644473 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2645
- Primary Citation Related Structures:
4QVB - PubMed Abstract:
Coenzyme F420 is a deazaflavin hydride carrier with a lower reduction potential than most flavins. In Mycobacterium tuberculosis (Mtb), F420 plays an important role in activating PA-824, an antituberculosis drug currently used in clinical trials. Although F420 is important to Mtb redox metabolism, little is known about the enzymes that bind F420 and the reactions that they catalyze. We have identified a novel F420 -binding protein, Rv1155, which is annotated in the Mtb genome sequence as a putative flavin mononucleotide (FMN)-binding protein. Using biophysical techniques, we have demonstrated that instead of binding FMN or other flavins, Rv1155 binds coenzyme F420 . The crystal structure of the complex of Rv1155 and F420 reveals one F420 molecule bound to each monomer of the Rv1155 dimer. Structural, biophysical, and bioinformatic analyses of the Rv1155-F420 complex provide clues about its role in the bacterium.
- Tuberculosis Research Section, National Institute of Allergy and Infectious Diseases, Bethesda, Maryland, 20892; Department of Chemistry, University of Cambridge, Cambridge, CB2 1EW, United Kingdom.
Organizational Affiliation: