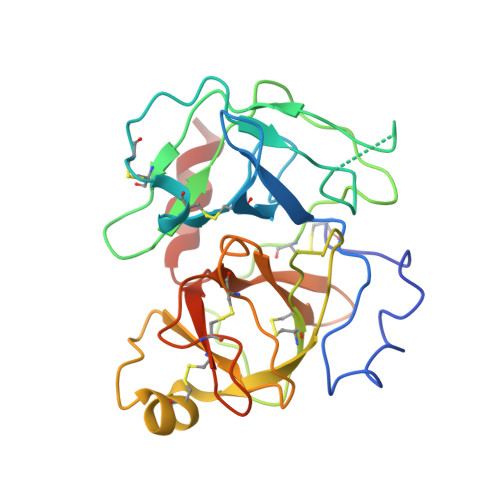
Structural basis for the binding specificity of human Recepteur d'Origine Nantais (RON) receptor tyrosine kinase to macrophage-stimulating protein.
Chao, K.L., Gorlatova, N.V., Eisenstein, E., Herzberg, O.(2014) J Biological Chem 289: 29948-29960
- PubMed: 25193665 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.594341
- Primary Citation Related Structures:
4QT8 - PubMed Abstract:
Recepteur d'origine nantais (RON) receptor tyrosine kinase and its ligand, serum macrophage-stimulating protein (MSP), play important roles in inflammation, cell growth, migration, and epithelial to mesenchymal transition during tumor development. The binding of mature MSPαβ (disulfide-linked α- and β-chains) to RON ectodomain modulates receptor dimerization, followed by autophosphorylation of tyrosines in the cytoplasmic receptor kinase domains. Receptor recognition is mediated by binding of MSP β-chain (MSPβ) to the RON Sema. Here we report the structure of RON Sema-PSI-IPT1 (SPI1) domains in complex with MSPβ at 3.0 Å resolution. The MSPβ serine protease-like β-barrel uses the degenerate serine protease active site to recognize blades 2, 3, and 4 of the β-propeller fold of RON Sema. Despite the sequence homology between RON and MET receptor tyrosine kinase and between MSP and hepatocyte growth factor, it is well established that there is no cross-reactivity between the two receptor-ligand systems. Comparison of the structure of RON SPI1 in complex with MSPβ and that of MET receptor tyrosine kinase Sema-PSI in complex with hepatocyte growth factor β-chain reveals the receptor-ligand selectivity determinants. Analytical ultracentrifugation studies of the SPI1-MSPβ interaction confirm the formation of a 1:1 complex. SPI1 and MSPαβ also associate primarily as a 1:1 complex with a binding affinity similar to that of SPI1-MSPβ. In addition, the SPI1-MSPαβ ultracentrifuge studies reveal a low abundance 2:2 complex with ∼ 10-fold lower binding affinity compared with the 1:1 species. These results support the hypothesis that the α-chain of MSPαβ mediates RON dimerization.
- From the Institute for Bioscience and Biotechnology Research, University of Maryland, Rockville, Maryland 20850 and.
Organizational Affiliation: