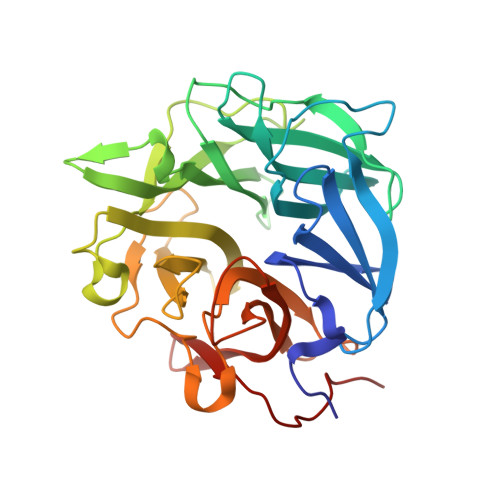
High-resolution crystal structure of a polyextreme GH43 glycosidase from Halothermothrix orenii with alpha-L-arabinofuranosidase activity.
Hassan, N., Kori, L.D., Gandini, R., Patel, B.K., Divne, C., Tan, T.C.(2015) Acta Crystallogr Sect F Struct Biol Cryst Commun 71: 338-345
- PubMed: 25760712 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X15003337
- Primary Citation Related Structures:
4QQS - PubMed Abstract:
A gene from the heterotrophic, halothermophilic marine bacterium Halothermothrix orenii has been cloned and overexpressed in Escherichia coli. This gene encodes the only glycoside hydrolase of family 43 (GH43) produced by H. orenii. The crystal structure of the H. orenii glycosidase was determined by molecular replacement and refined at 1.10 Å resolution. As for other GH43 members, the enzyme folds as a five-bladed β-propeller. The structure features a metal-binding site on the propeller axis, near the active site. Based on thermal denaturation data, the H. orenii glycosidase depends on divalent cations in combination with high salt for optimal thermal stability against unfolding. A maximum melting temperature of 76°C was observed in the presence of 4 M NaCl and Mn(2+) at pH 6.5. The gene encoding the H. orenii GH43 enzyme has previously been annotated as a putative α-L-arabinofuranosidase. Activity was detected with p-nitrophenyl-α-L-arabinofuranoside as a substrate, and therefore the name HoAraf43 was suggested for the enzyme. In agreement with the conditions for optimal thermal stability against unfolding, the highest arabinofuranosidase activity was obtained in the presence of 4 M NaCl and Mn(2+) at pH 6.5, giving a specific activity of 20-36 µmol min(-1) mg(-1). The active site is structurally distinct from those of other GH43 members, including arabinanases, arabinofuranosidases and xylanases. This probably reflects the special requirements for degrading the unique biomass available in highly saline aqueous ecosystems, such as halophilic algae and halophytes. The amino-acid distribution of HoAraf43 has similarities to those of mesophiles, thermophiles and halophiles, but also has unique features, for example more hydrophobic amino acids on the surface and fewer buried charged residues.
- School of Biotechnology, KTH Royal Institute of Technology, Stockholm, Sweden.
Organizational Affiliation: