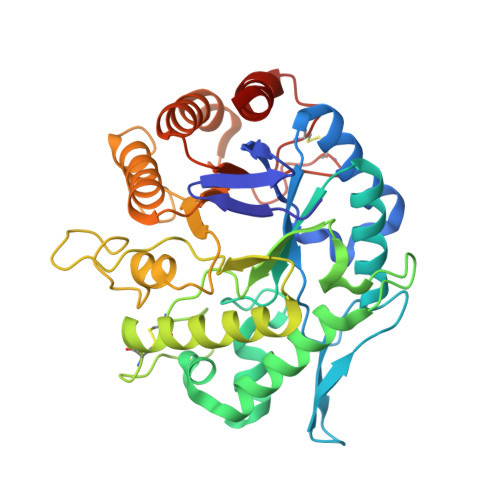
Structural insights into the substrate specificity and transglycosylation activity of a fungal glycoside hydrolase family 5 beta-mannosidase.
Zhou, P., Liu, Y., Yan, Q., Chen, Z., Qin, Z., Jiang, Z.(2014) Acta Crystallogr D Biol Crystallogr 70: 2970-2982
- PubMed: 25372687 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004714019762
- Primary Citation Related Structures:
4LYP, 4LYQ, 4LYR, 4NRR, 4NRS, 4QP0 - PubMed Abstract:
β-Mannosidases are exo-acting glycoside hydrolases (GHs) that catalyse the removal of the nonreducing end β-D-mannose from manno-oligosaccharides or mannoside-substituted molecules. They play important roles in fundamental biological processes and also have potential applications in various industries. In this study, the first fungal GH family 5 β-mannosidase (RmMan5B) from Rhizomucor miehei was functionally and structurally characterized. RmMan5B exhibited a much higher activity against manno-oligosaccharides than against p-nitrophenyl β-D-mannopyranoside (pNPM) and had a transglycosylation activity which transferred mannosyl residues to sugars such as fructose. To investigate its substrate specificity and transglycosylation activity, crystal structures of RmMan5B and of its inactive E202A mutant in complex with mannobiose, mannotriose and mannosyl-fructose were determined at resolutions of 1.3, 2.6, 2.0 and 2.4 Å, respectively. In addition, the crystal structure of R. miehei β-mannanase (RmMan5A) was determined at a resolution of 2.3 Å. Both RmMan5A and RmMan5B adopt the (β/α)8-barrel architecture, which is globally similar to the other members of GH family 5. However, RmMan5B shows several differences in the loop around the active site. The extended loop between strand β8 and helix α8 (residues 354-392) forms a `double' steric barrier to `block' the substrate-binding cleft at the end of the -1 subsite. Trp119, Asn260 and Glu380 in the β-mannosidase, which are involved in hydrogen-bond contacts with the -1 mannose, might be essential for exo catalytic activity. Moreover, the structure of RmMan5B in complex with mannosyl-fructose has provided evidence for the interactions between the β-mannosidase and D-fructofuranose. Overall, the present study not only helps in understanding the catalytic mechanism of GH family 5 β-mannosidases, but also provides a basis for further enzymatic engineering of β-mannosidases and β-mannanases.
- Department of Biotechnology, College of Food Science and Nutritional Engineering, China Agricultural University, Beijing 100083, People's Republic of China.
Organizational Affiliation: