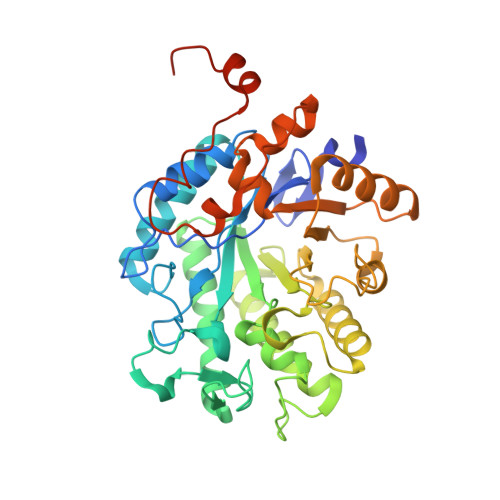
Structure of an Aspergillus fumigatus old yellow enzyme (EasA) involved in ergot alkaloid biosynthesis.
Chilton, A.S., Ellis, A.L., Lamb, A.L.(2014) Acta Crystallogr Sect F Struct Biol Cryst Commun 70: 1328-1332
- PubMed: 25286934 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14018962
- Primary Citation Related Structures:
4QNW - PubMed Abstract:
The Aspergillus fumigatus old yellow enzyme (OYE) EasA reduces chanoclavine-I aldehyde to dihydrochanoclavine aldehyde and works in conjunction with festuclavine synthase at the branchpoint for ergot alkaloid pathways. The crystal structure of the FMN-loaded EasA was determined to 1.8 Å resolution. The active-site amino acids of OYE are conserved, supporting a similar mechanism for reduction of the α/β-unsaturated aldehyde. The C-terminal tail of one monomer packs into the active site of a monomer in the next asymmetric unit, which is most likely to be a crystallization artifact and not a mechanism of self-regulation.
- Molecular Biosciences, University of Kansas, 1200 Sunnyside Avenue, Lawrence, KS 66045, USA.
Organizational Affiliation: