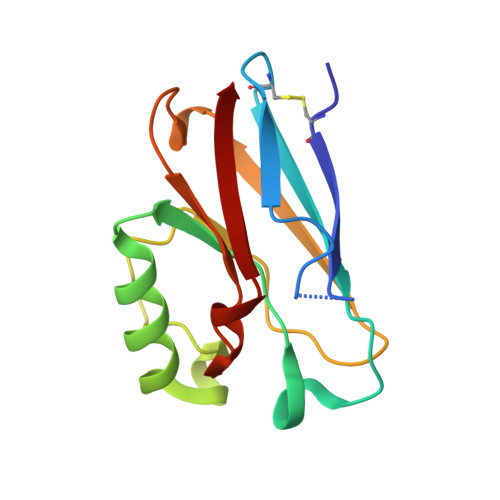
Redesigning the Blue Copper Azurin into a Redox-Active Mononuclear Nonheme Iron Protein: Preparation and Study of Fe(II)-M121E Azurin.
Liu, J., Meier, K.K., Tian, S., Zhang, J.L., Guo, H., Schulz, C.E., Robinson, H., Nilges, M.J., Munck, E., Lu, Y.(2014) J Am Chem Soc 136: 12337-12344
- PubMed: 25082811 Search on PubMed
- DOI: https://doi.org/10.1021/ja505410u
- Primary Citation Related Structures:
4QKT, 4QLW - PubMed Abstract:
Much progress has been made in designing heme and dinuclear nonheme iron enzymes. In contrast, engineering mononuclear nonheme iron enzymes is lagging, even though these enzymes belong to a large class that catalyzes quite diverse reactions. Herein we report spectroscopic and X-ray crystallographic studies of Fe(II)-M121E azurin (Az), by replacing the axial Met121 and Cu(II) in wild-type azurin (wtAz) with Glu and Fe(II), respectively. In contrast to the redox inactive Fe(II)-wtAz, the Fe(II)-M121EAz mutant can be readily oxidized by Na2IrCl6, and interestingly, the protein exhibits superoxide scavenging activity. Mössbauer and EPR spectroscopies, along with X-ray structural comparisons, revealed similarities and differences between Fe(II)-M121EAz, Fe(II)-wtAz, and superoxide reductase (SOR) and allowed design of the second generation mutant, Fe(II)-M121EM44KAz, that exhibits increased superoxide scavenging activity by 2 orders of magnitude. This finding demonstrates the importance of noncovalent secondary coordination sphere interactions in fine-tuning enzymatic activity.
- Department of Chemistry, University of Illinois at Urbana-Champaign , Urbana, Illinois 61801, United States.
Organizational Affiliation: