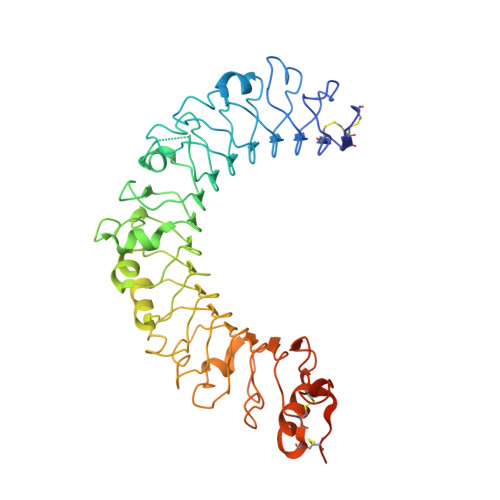
Crystal structure of the C-terminal domain of mouse TLR9.
Collins, B., Wilson, I.A.(2014) Proteins 82: 2874-2878
- PubMed: 24888966 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.24616
- Primary Citation Related Structures:
4QDH - PubMed Abstract:
Toll-like receptors (TLRs) are important pattern recognition receptors that function in innate immunity. Elucidating the structure and signaling mechanisms of TLR9, a sensor of foreign and endogenous DNA, is essential for understanding its key role in immunity against microbial pathogens as well as in autoimmunity. Abundant evidence suggests that the TLR9-CTD (C-terminal domain) by itself is capable of DNA binding and signaling. The crystal structure of unliganded mouse TLR9-CTD is presented. TLR9-CTD exhibits one unique feature, a cluster of stacked aromatic and arginine side chains on its concave face. Overall, its structure is most related to the TLR8-CTD, suggesting a similar mode of ligand binding and signaling.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, California.
Organizational Affiliation: