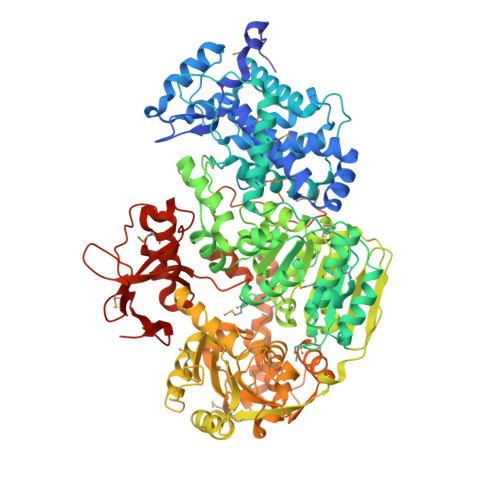
Molecular insights into DNA interference by CRISPR-associated nuclease-helicase Cas3.
Gong, B., Shin, M., Sun, J., Jung, C.H., Bolt, E.L., van der Oost, J., Kim, J.S.(2014) Proc Natl Acad Sci U S A 111: 16359-16364
- PubMed: 25368186 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1410806111
- Primary Citation Related Structures:
4Q2C, 4Q2D - PubMed Abstract:
Mobile genetic elements in bacteria are neutralized by a system based on clustered regularly interspaced short palindromic repeats (CRISPRs) and CRISPR-associated (Cas) proteins. Type I CRISPR-Cas systems use a "Cascade" ribonucleoprotein complex to guide RNA specifically to complementary sequence in invader double-stranded DNA (dsDNA), a process called "interference." After target recognition by Cascade, formation of an R-loop triggers recruitment of a Cas3 nuclease-helicase, completing the interference process by destroying the invader dsDNA. To elucidate the molecular mechanism of CRISPR interference, we analyzed crystal structures of Cas3 from the bacterium Thermobaculum terrenum, with and without a bound ATP analog. The structures reveal a histidine-aspartate (HD)-type nuclease domain fused to superfamily-2 (SF2) helicase domains and a distinct C-terminal domain. Binding of ATP analog at the interface of the SF2 helicase RecA-like domains rearranges a motif V with implications for the enzyme mechanism. The HD-nucleolytic site contains two metal ions that are positioned at the end of a proposed nucleic acid-binding tunnel running through the SF2 helicase structure. This structural alignment suggests a mechanism for 3' to 5' nucleolytic processing of the displaced strand of invader DNA that is coordinated with ATP-dependent 3' to 5' translocation of Cas3 along DNA. In agreement with biochemical studies, the presented Cas3 structures reveal important mechanistic details on the neutralization of genetic invaders by type I CRISPR-Cas systems.
- Interdisciplinary Graduate Program in Molecular Medicine, Chonnam National University, Gwangju 501-746, Korea;
Organizational Affiliation: