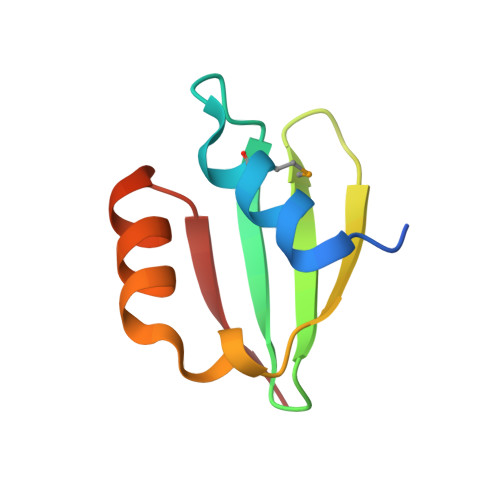
Structures of periplasmic and transmembrane domains of FtsH suggest a reverse translocon mechanism for protein extraction from membrane
An, J.Y., Sharif, H., Barrera, F.N., Karabadzhak, A., Kang, G.B., Park, K.J., Sakkiah, S., Lee, K.W., Lee, S., Engelman, D.M., Wang, J., Eom, S.H.To be published.