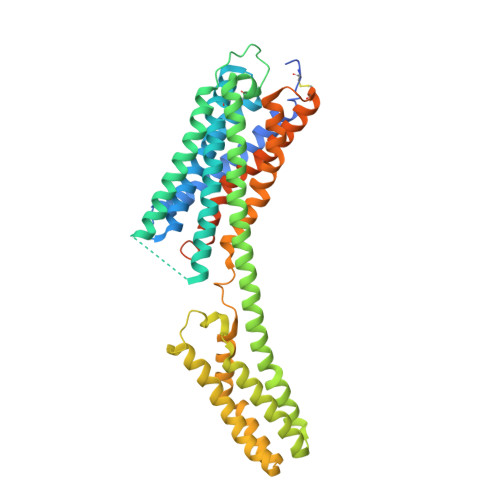
Agonist-bound structure of the human P2Y12 receptor
Zhang, J., Zhang, K., Gao, Z.G., Paoletta, S., Zhang, D., Han, G.W., Li, T., Ma, L., Zhang, W., Muller, C.E., Yang, H., Jiang, H., Cherezov, V., Katritch, V., Jacobson, K.A., Stevens, R.C., Wu, B., Zhao, Q.(2014) Nature 509: 119-122
- PubMed: 24784220 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature13288
- Primary Citation Related Structures:
4PXZ, 4PY0 - PubMed Abstract:
The P2Y12 receptor (P2Y12R), one of eight members of the P2YR family expressed in humans, is one of the most prominent clinical drug targets for inhibition of platelet aggregation. Although mutagenesis and modelling studies of the P2Y12R provided useful insights into ligand binding, the agonist and antagonist recognition and function at the P2Y12R remain poorly understood at the molecular level. Here we report the structures of the human P2Y12R in complex with the full agonist 2-methylthio-adenosine-5'-diphosphate (2MeSADP, a close analogue of endogenous agonist ADP) at 2.5 Å resolution, and the corresponding ATP derivative 2-methylthio-adenosine-5'-triphosphate (2MeSATP) at 3.1 Å resolution. These structures, together with the structure of the P2Y12R with antagonist ethyl 6-(4-((benzylsulfonyl)carbamoyl)piperidin-1-yl)-5-cyano-2-methylnicotinate (AZD1283), reveal striking conformational changes between nucleotide and non-nucleotide ligand complexes in the extracellular regions. Further analysis of these changes provides insight into a distinct ligand binding landscape in the δ-group of class A G-protein-coupled receptors (GPCRs). Agonist and non-nucleotide antagonist adopt different orientations in the P2Y12R, with only partially overlapped binding pockets. The agonist-bound P2Y12R structure answers long-standing questions surrounding P2Y12R-agonist recognition, and reveals interactions with several residues that had not been reported to be involved in agonist binding. As a first example, to our knowledge, of a GPCR in which agonist access to the binding pocket requires large-scale rearrangements in the highly malleable extracellular region, the structural and docking studies will therefore provide invaluable insight into the pharmacology and mechanisms of action of agonists and different classes of antagonists for the P2Y12R and potentially for other closely related P2YRs.
- 1] CAS Key Laboratory of Receptor Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, 555 Zuchongzhi Road, Pudong, Shanghai 201203, China [2].
Organizational Affiliation: