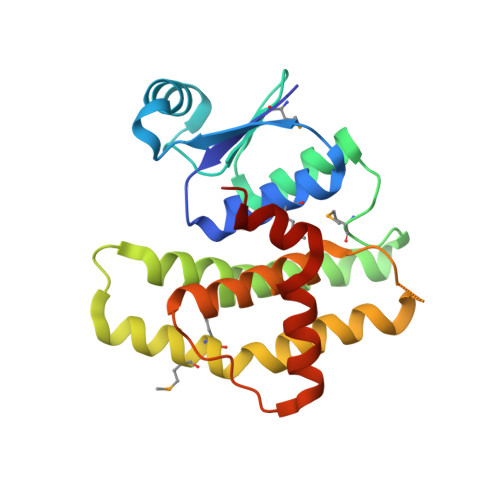
Structural and Biochemical Characterization of the Francisella tularensis Pathogenicity Regulator, Macrophage Locus Protein A (MglA).
Cuthbert, B.J., Brennan, R.G., Schumacher, M.A.(2015) PLoS One 10: e0128225-e0128225
- PubMed: 26121147 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0128225
- Primary Citation Related Structures:
4PUR - PubMed Abstract:
Francisella tularensis is one of the most infectious bacteria known and is the etiologic agent of tularemia. Francisella virulence arises from a 33 kilobase (Kb) pathogenicity island (FPI) that is regulated by the macrophage locus protein A (MglA) and the stringent starvation protein A (SspA). These proteins interact with both RNA polymerase (RNAP) and the pathogenicity island gene regulator (PigR) to activate FPI transcription. However, the molecular mechanisms involved are not well understood. Indeed, while most bacterial SspA proteins function as homodimers to activate transcription, F. tularensis SspA forms a heterodimer with the MglA protein, which is unique to F. tularensis. To gain insight into MglA function, we performed structural and biochemical studies. The MglA structure revealed that it contains a fold similar to the SspA protein family. Unexpectedly, MglA also formed a homodimer in the crystal. Chemical crosslinking and size exclusion chromatography (SEC) studies showed that MglA is able to self-associate in solution to form a dimer but that it preferentially heterodimerizes with SspA. Finally, the MglA structure revealed malate, which was used in crystallization, bound in an open pocket formed by the dimer, suggesting the possibility that this cleft could function in small molecule ligand binding. The location of this binding region relative to recently mapped PigR and RNAP interacting sites suggest possible roles for small molecule binding in MglA and SspA•MglA function.
- Department of Biochemistry, Duke University School of Medicine, Durham, North Carolina, 27710, United States of America.
Organizational Affiliation: