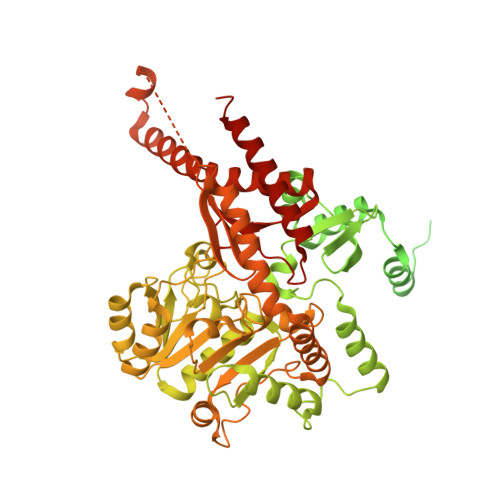
Crystal structure of the catalytic domain of PigE: a transaminase involved in the biosynthesis of 2-methyl-3-n-amyl-pyrrole (MAP) from Serratia sp. FS14
Lou, X.D., Ran, T.T., Han, N., Gao, Y., He, J., Tang, L., Xu, D.Q., Wang, W.W.(2014) Biochem Biophys Res Commun 447: 178-183
- PubMed: 24704447 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2014.03.125
- Primary Citation Related Structures:
4PPM - PubMed Abstract:
Prodigiosin, a tripyrrole red pigment synthesized by Serratia and some other microbes through a bifurcated biosynthesis pathway, MBC (4-methoxy-2,2'-bipyrrole-5-carbaldehyde) and MAP (2-methyl-3-n-amyl-pyrrole) are synthesized separately and then condensed by PigC to form prodigiosin. MAP is synthesized sequentially by PigD, PigE and PigB. PigE catalyzes the transamination of an amino group to the aldehyde group of 3-acetyloctanal, resulting in an aminoketone, which spontaneously cyclizes to form H2MAP. Here we report the crystal structure of the catalytic domain of PigE which involved in the biosynthesis of prodigiosin precursor MAP for the first time to a resolution of 2.3Å with a homodimer in the asymmetric unit. The monomer of PigE catalytic domain is composed of three domains with PLP as cofactor: a small N-terminal domain connecting the catalytic domain with the front part of PigE, a large PLP-binding domain and a C-terminal domain. The residues from both monomers build the PLP binding site at the interface of the dimer which resembles the other PLP-dependent enzymes. Structural comparison of PigE with Thermus thermophilus AcOAT showed a higher hydrophobic and smaller active site of PigE, these differences may be the reason for substrate specificity.
- Key Laboratory of Agricultural and Environmental Microbiology, Ministry of Agriculture, College of Life Sciences, Nanjing Agricultural University, 210095 Nanjing, China.
Organizational Affiliation: