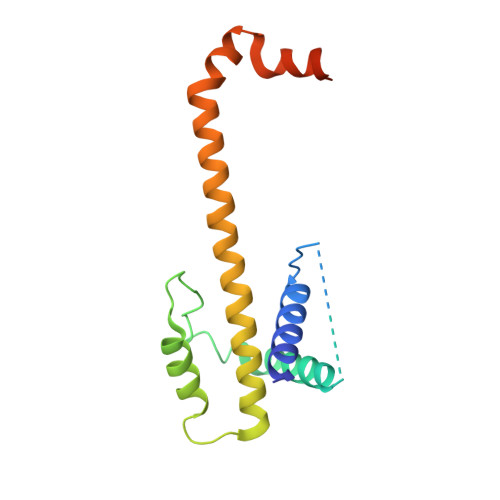
Structural transition in Bcl-xL and its potential association with mitochondrial calcium ion transport
Rajan, S., Choi, M., Nguyen, Q.T., Ye, H., Liu, W., Toh, H.T., Kang, C., Kamariah, N., Li, C., Huang, H., White, C., Baek, K., Gruber, G., Yoon, H.S.(2015) Sci Rep 5: 10609-10609
- PubMed: 26023881 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep10609
- Primary Citation Related Structures:
4PPI - PubMed Abstract:
Bcl-2 family proteins are key regulators for cellular homeostasis in response to apoptotic stimuli. Bcl-xL, an antiapoptotic Bcl-2 family member, undergoes conformational transitions, which leads to two conformational states: the cytoplasmic and membrane-bound. Here we present the crystal and small-angle X-ray scattering (SAXS) structures of Bcl-xL treated with the mild detergent n-Octyl β-D-Maltoside (OM). The detergent-treated Bcl-xL forms a dimer through three-dimensional domain swapping (3DDS) by swapping helices α6-α8 between two monomers. Unlike Bax, a proapoptotic member of the Bcl-2 family, Bcl-xL is not converted to 3DDS homodimer upon binding BH3 peptides and ABT-737, a BH3 mimetic drug. We also designed Bcl-xL mutants which cannot dimerize and show that these mutants reduced mitochondrial calcium uptake in MEF cells. This illustrates the structural plasticity in Bcl-xL providing hints toward the probable molecular mechanism for Bcl-xL to play a regulatory role in mitochondrial calcium ion transport.
- School of Biological Sciences, Nanyang Technological University, 60 Nanyang Drive, Singapore 637551.
Organizational Affiliation: