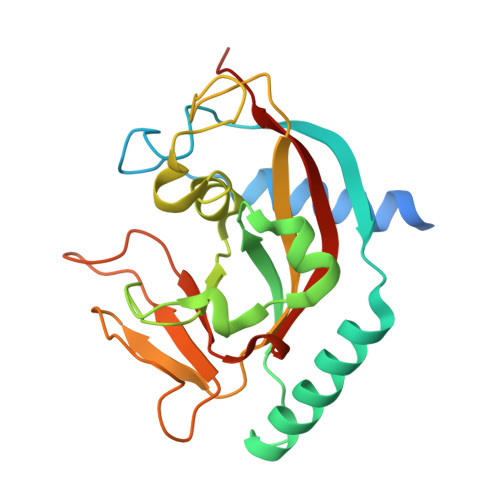
Insights into the binding of PARP inhibitors to the catalytic domain of human tankyrase-2.
Qiu, W., Lam, R., Voytyuk, O., Romanov, V., Gordon, R., Gebremeskel, S., Vodsedalek, J., Thompson, C., Beletskaya, I., Battaile, K.P., Pai, E.F., Rottapel, R., Chirgadze, N.Y.(2014) Acta Crystallogr D Biol Crystallogr 70: 2740-2753
- PubMed: 25286857 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004714017660
- Primary Citation Related Structures:
4PML, 4PNL, 4PNM, 4PNN, 4PNQ, 4PNR, 4PNS, 4PNT, 4TJU, 4TJW, 4TJY, 4TK0, 4TK5, 4TKF, 4TKG, 4TKI - PubMed Abstract:
The poly(ADP-ribose) polymerase (PARP) family represents a new class of therapeutic targets with diverse potential disease indications. PARP1 and PARP2 inhibitors have been developed for breast and ovarian tumors manifesting double-stranded DNA-repair defects, whereas tankyrase 1 and 2 (TNKS1 and TNKS2, also known as PARP5a and PARP5b, respectively) inhibitors have been developed for tumors with elevated β-catenin activity. As the clinical relevance of PARP inhibitors continues to be actively explored, there is heightened interest in the design of selective inhibitors based on the detailed structural features of how small-molecule inhibitors bind to each of the PARP family members. Here, the high-resolution crystal structures of the human TNKS2 PARP domain in complex with 16 various PARP inhibitors are reported, including the compounds BSI-201, AZD-2281 and ABT-888, which are currently in Phase 2 or 3 clinical trials. These structures provide insight into the inhibitor-binding modes for the tankyrase PARP domain and valuable information to guide the rational design of future tankyrase-specific inhibitors.
- Princess Margaret Cancer Center, University Health Network, Toronto, Ontario, Canada.
Organizational Affiliation: