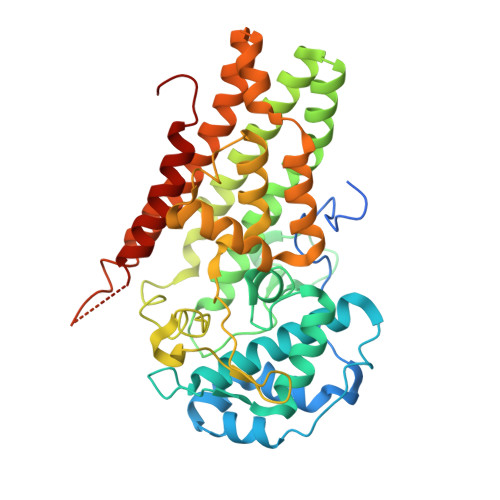
Crystal Structures and Structure-Activity Relationships of Imidazothiazole Derivatives as IDO1 Inhibitors.
Tojo, S., Kohno, T., Tanaka, T., Kamioka, S., Ota, Y., Ishii, T., Kamimoto, K., Asano, S., Isobe, Y.(2014) ACS Med Chem Lett 5: 1119-1123
- PubMed: 25313323 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml500247w
- Primary Citation Related Structures:
4PK5, 4PK6 - PubMed Abstract:
Indoleamine 2,3-dioxygenase 1 (IDO1) is considered as a promising target for the treatment of several diseases, including neurological disorders and cancer. We report here the crystal structures of two IDO1/IDO1 inhibitor complexes, one of which shows that Amg-1 is directly bound to the heme iron of IDO1 with a clear induced fit. We also describe the identification and preliminary optimization of imidazothiazole derivatives as novel IDO1 inhibitors. Using our crystal structure information and structure-activity relationships (SAR) at the pocket-B of IDO1, we found a series of urea derivatives as potent IDO1 inhibitors and revealed that generation of an induced fit and the resulting interaction with Phe226 and Arg231 are essential for potent IDO1 inhibitory activity. The results of this study are very valuable for understanding the mechanism of IDO1 activation, which is very important for structure-based drug design (SBDD) to discover potent IDO1 inhibitors.
- Dainippon Sumitomo Pharma Co., Ltd. , 3-1-98 Kasugade-naka, Konohana-ku, Osaka 554-0022, Japan.
Organizational Affiliation: