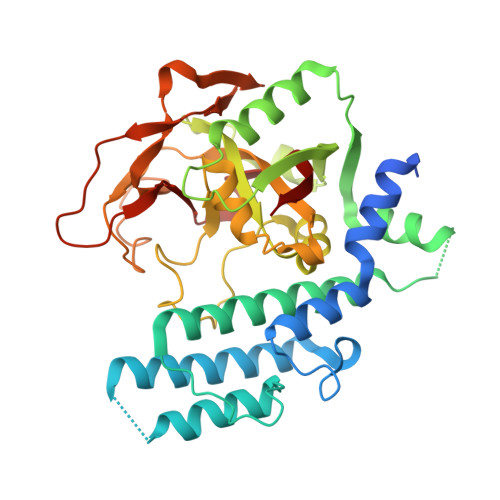
Structural basis for the inhibition of poly(ADP-ribose) polymerases 1 and 2 by BMN 673, a potent inhibitor derived from dihydropyridophthalazinone.
Aoyagi-Scharber, M., Gardberg, A.S., Yip, B.K., Wang, B., Shen, Y., Fitzpatrick, P.A.(2014) Acta Crystallogr F Struct Biol Commun 70: 1143-1149
- PubMed: 25195882 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14015088
- Primary Citation Related Structures:
4PJT, 4PJV - PubMed Abstract:
Poly(ADP-ribose) polymerases 1 and 2 (PARP1 and PARP2), which are involved in DNA damage response, are targets of anticancer therapeutics. BMN 673 is a novel PARP1/2 inhibitor with substantially increased PARP-mediated tumor cytotoxicity and is now in later-stage clinical development for BRCA-deficient breast cancers. In co-crystal structures, BMN 673 is anchored to the nicotinamide-binding pocket via an extensive network of hydrogen-bonding and π-stacking interactions, including those mediated by active-site water molecules. The novel di-branched scaffold of BMN 673 extends the binding interactions towards the outer edges of the pocket, which exhibit the least sequence homology among PARP enzymes. The crystallographic structural analyses reported here therefore not only provide critical insights into the molecular basis for the exceptionally high potency of the clinical development candidate BMN 673, but also new opportunities for increasing inhibitor selectivity.
- Research and Drug Discovery, BioMarin Pharmaceutical Inc., 105 Digital Drive, Novato, CA 94949, USA.
Organizational Affiliation: