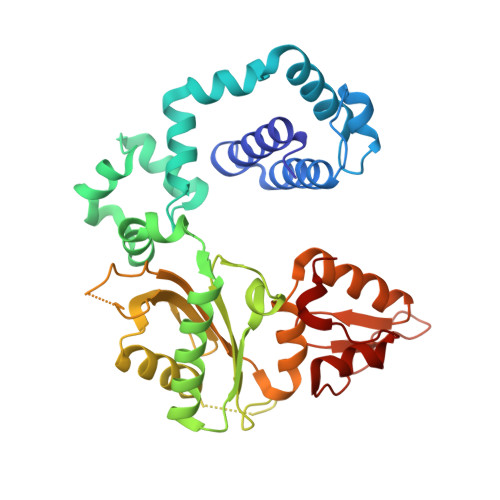
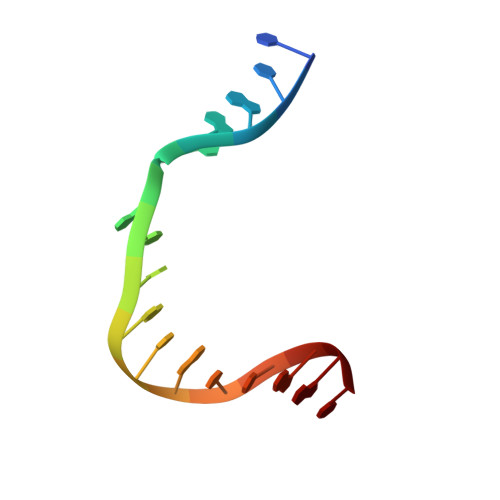
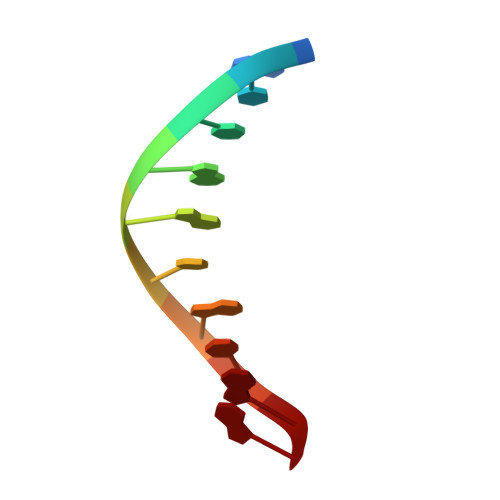
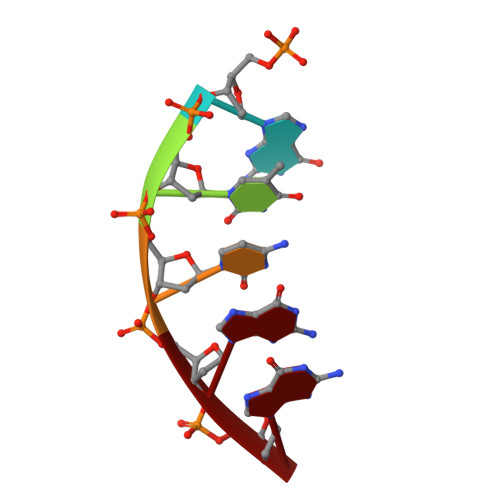
The spontaneous replication error and the mismatch discrimination mechanisms of human DNA polymerase beta.
Koag, M.C., Nam, K., Lee, S.(2015) Nucleic Acids Res 42: 11233-11245
- PubMed: 25200079 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gku789
- Primary Citation Related Structures:
4PGQ, 4PGX, 4PHA, 4PHD - PubMed Abstract:
To provide molecular-level insights into the spontaneous replication error and the mismatch discrimination mechanisms of human DNA polymerase β (polβ), we report four crystal structures of polβ complexed with dG•dTTP and dA•dCTP mismatches in the presence of Mg2+ or Mn2+. The Mg(2+)-bound ground-state structures show that the dA•dCTP-Mg2+ complex adopts an 'intermediate' protein conformation while the dG•dTTP-Mg2+ complex adopts an open protein conformation. The Mn(2+)-bound 'pre-chemistry-state' structures show that the dA•dCTP-Mn2+ complex is structurally very similar to the dA•dCTP-Mg2+ complex, whereas the dG•dTTP-Mn2+ complex undergoes a large-scale conformational change to adopt a Watson-Crick-like dG•dTTP base pair and a closed protein conformation. These structural differences, together with our molecular dynamics simulation studies, suggest that polβ increases replication fidelity via a two-stage mismatch discrimination mechanism, where one is in the ground state and the other in the closed conformation state. In the closed conformation state, polβ appears to allow only a Watson-Crick-like conformation for purine•pyrimidine base pairs, thereby discriminating the mismatched base pairs based on their ability to form the Watson-Crick-like conformation. Overall, the present studies provide new insights into the spontaneous replication error and the replication fidelity mechanisms of polβ.
- Division of Medicinal Chemistry, College of Pharmacy, The University of Texas at Austin, Austin, TX 78712, USA.
Organizational Affiliation: