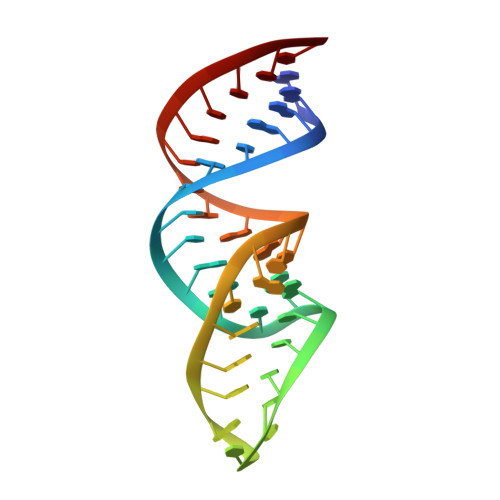
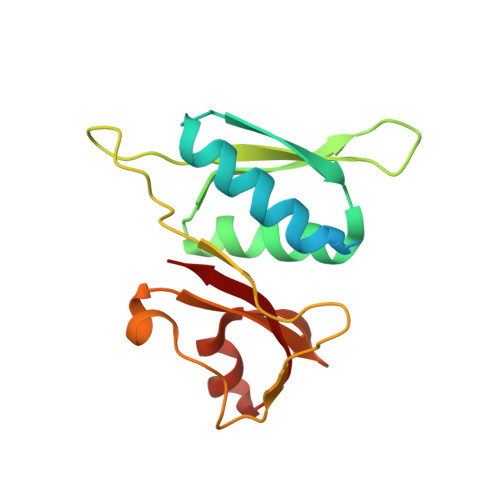
Structure analysis of free and bound states of an RNA aptamer against ribosomal protein S8 from Bacillus anthracis.
Davlieva, M., Donarski, J., Wang, J., Shamoo, Y., Nikonowicz, E.P.(2014) Nucleic Acids Res 42: 10795-10808
- PubMed: 25140011 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gku743
- Primary Citation Related Structures:
4PDB - PubMed Abstract:
Several protein-targeted RNA aptamers have been identified for a variety of applications and although the affinities of numerous protein-aptamer complexes have been determined, the structural details of these complexes have not been widely explored. We examined the structural accommodation of an RNA aptamer that binds bacterial r-protein S8. The core of the primary binding site for S8 on helix 21 of 16S rRNA contains a pair of conserved base triples that mold the sugar-phosphate backbone to S8. The aptamer, which does not contain the conserved sequence motif, is specific for the rRNA binding site of S8. The protein-free RNA aptamer adopts a helical structure with multiple non-canonical base pairs. Surprisingly, binding of S8 leads to a dramatic change in the RNA conformation that restores the signature S8 recognition fold through a novel combination of nucleobase interactions. Nucleotides within the non-canonical core rearrange to create a G-(G-C) triple and a U-(A-U)-U quartet. Although native-like S8-RNA interactions are present in the aptamer-S8 complex, the topology of the aptamer RNA differs from that of the helix 21-S8 complex. This is the first example of an RNA aptamer that adopts substantially different secondary structures in the free and protein-bound states and highlights the remarkable plasticity of RNA secondary structure.
- Department of Biochemistry and Cell Biology, Rice University, Houston, TX 77251-1892, USA.
Organizational Affiliation: