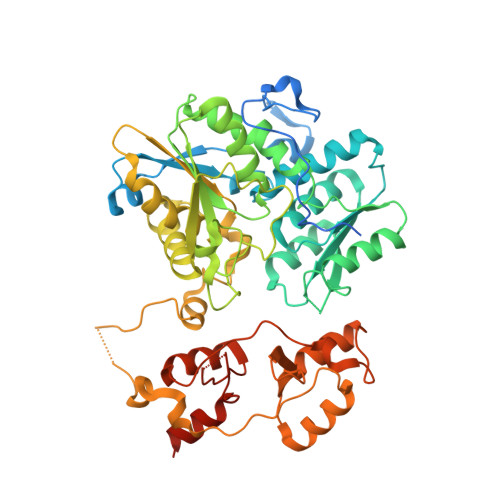
Structural insight into the molecular mechanism of allosteric activation of human cystathionine beta-synthase by S-adenosylmethionine.
Ereno-Orbea, J., Majtan, T., Oyenarte, I., Kraus, J.P., Martinez-Cruz, L.A.(2014) Proc Natl Acad Sci U S A 111: E3845-E3852
- PubMed: 25197074
- DOI: https://doi.org/10.1073/pnas.1414545111
- Primary Citation of Related Structures:
4PCU - PubMed Abstract:
Cystathionine β-synthase (CBS) is a heme-dependent and pyridoxal-5'-phosphate-dependent protein that controls the flux of sulfur from methionine to cysteine, a precursor of glutathione, taurine, and H2S. Deficiency of CBS activity causes homocystinuria, the most frequent disorder of sulfur amino acid metabolism. In contrast to CBSs from lower organisms, human CBS (hCBS) is allosterically activated by S-adenosylmethionine (AdoMet), which binds to the regulatory domain and triggers a conformational change that allows the protein to progress from the basal toward the activated state. The structural basis of the underlying molecular mechanism has remained elusive so far. Here, we present the structure of hCBS with bound AdoMet, revealing the activated conformation of the human enzyme. Binding of AdoMet triggers a conformational change in the Bateman module of the regulatory domain that favors its association with a Bateman module of the complementary subunit to form an antiparallel CBS module. Such an arrangement is very similar to that found in the constitutively activated insect CBS. In the presence of AdoMet, the autoinhibition exerted by the regulatory region is eliminated, allowing for improved access of substrates to the catalytic pocket. Based on the availability of both the basal and the activated structures, we discuss the mechanism of hCBS activation by AdoMet and the properties of the AdoMet binding site, as well as the responsiveness of the enzyme to its allosteric regulator. The structure described herein paves the way for the rational design of compounds modulating hCBS activity and thus transsulfuration, redox status, and H2S biogenesis.
- Structural Biology Unit, Center for Cooperative Research in Biosciences (CIC bioGUNE), Technology Park of Bizkaia, 48160 Derio, Spain; and.
Organizational Affiliation: