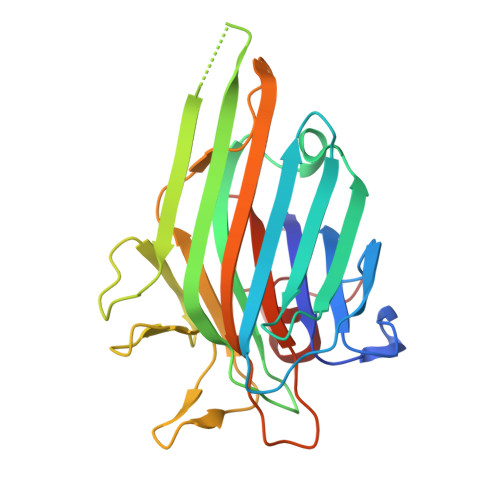
Crystal structure of Canavalia brasiliensis seed lectin (ConBr) complexed with Gamma-Aminobutyric Acid (GABA)
Delatorre, P., Rocha, B.A.M., Silva-Filho, J.C., Teixeira, C.S., Cavada, B.S., Nascimento, K.S., Nbrega, R.B., Nagano, C.S., Sampaio, A.H., Leal, R.B., Neto, I.L.B.To be published.