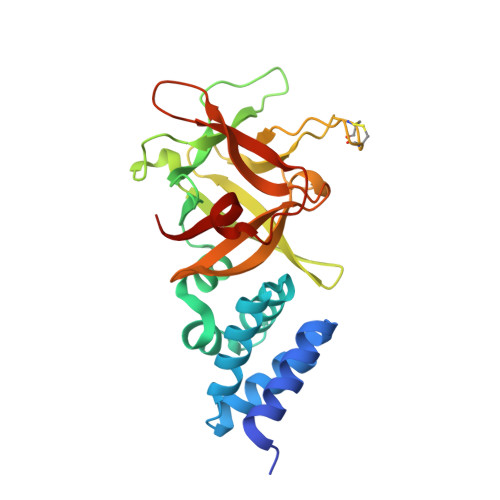
Crystal structure of Bombyx mori lipoprotein 6: comparative structural analysis of the 30-kDa lipoprotein family.
Pietrzyk, A.J., Bujacz, A., ochynska, M., Jaskolski, M., Bujacz, G.(2014) PLoS One 9: e108761-e108761
- PubMed: 25379889 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0108761
- Primary Citation Related Structures:
4PC4 - PubMed Abstract:
The 30-kDa lipoprotein (LP) family of mulberry silkworm comprises major hemolymph proteins specific to the fifth instar larvae. The family consists of 46 members, 24 of which are referred to as typical 30-kDa LPs. To date, two crystal structures of 30-kDa LPs from Bombyx mori have been described (Bmlp3 and Bmlp7). Here, we present the crystal structure of Bmlp6, another 30-kDa LP member. Bmlp6 is comprised of two domains characteristic of this family, the VHS-type N-terminal domain and β-trefoil C-terminal domain. The structures of the three 30-kDa LPs have been compared and a number of differences are noted, including loop conformation, the surface electrostatic potential, and the potential binding cavities. We discuss the observed structural differences in the light of the potential different roles of the particular 30-kDa LP members in silkworm physiology.
- Center for Biocrystallographic Research, Institute of Bioorganic Chemistry, Polish Academy of Sciences, Poznan, Poland.
Organizational Affiliation: