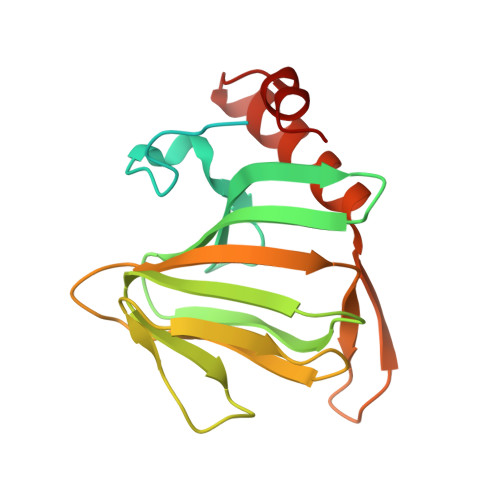
Structure of the 2,4'-dihydroxyacetophenone dioxygenase from Alcaligenes sp. 4HAP
Keegan, R., Lebedev, A., Erskine, P., Guo, J., Wood, S.P., Hopper, D.J., Rigby, S.E.J., Cooper, J.B.(2014) Acta Crystallogr D Biol Crystallogr 70: 2444-2454
- PubMed: 25195757 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004714015053
- Primary Citation Related Structures:
4P9G - PubMed Abstract:
The enzyme 2,4'-dihydroxyacetophenone dioxygenase (DAD) catalyses the conversion of 2,4'-dihydroxyacetophenone to 4-hydroxybenzoic acid and formic acid with the incorporation of molecular oxygen. Whilst the vast majority of dioxygenases cleave within the aromatic ring of the substrate, DAD is very unusual in that it is involved in C-C bond cleavage in a substituent of the aromatic ring. There is evidence that the enzyme is a homotetramer of 20.3 kDa subunits, each containing nonhaem iron, and its sequence suggests that it belongs to the cupin family of dioxygenases. In this paper, the first X-ray structure of a DAD enzyme from the Gram-negative bacterium Alcaligenes sp. 4HAP is reported, at a resolution of 2.2 Å. The structure establishes that the enzyme adopts a cupin fold, forming dimers with a pronounced hydrophobic interface between the monomers. The catalytic iron is coordinated by three histidine residues (76, 78 and 114) within a buried active-site cavity. The iron also appears to be tightly coordinated by an additional ligand which was putatively assigned as a carbonate dianion since this fits the electron density optimally, although it might also be the product formate. The modelled carbonate is located in a position which is highly likely to be occupied by the α-hydroxyketone group of the bound substrate during catalysis. Modelling of a substrate molecule in this position indicates that it will interact with many conserved amino acids in the predominantly hydrophobic active-site pocket where it undergoes peroxide radical-mediated heterolysis.
- STFC Rutherford Appleton Laboratory, RAL, Harwell Oxford, Didcot OX11 0FA, England.
Organizational Affiliation: